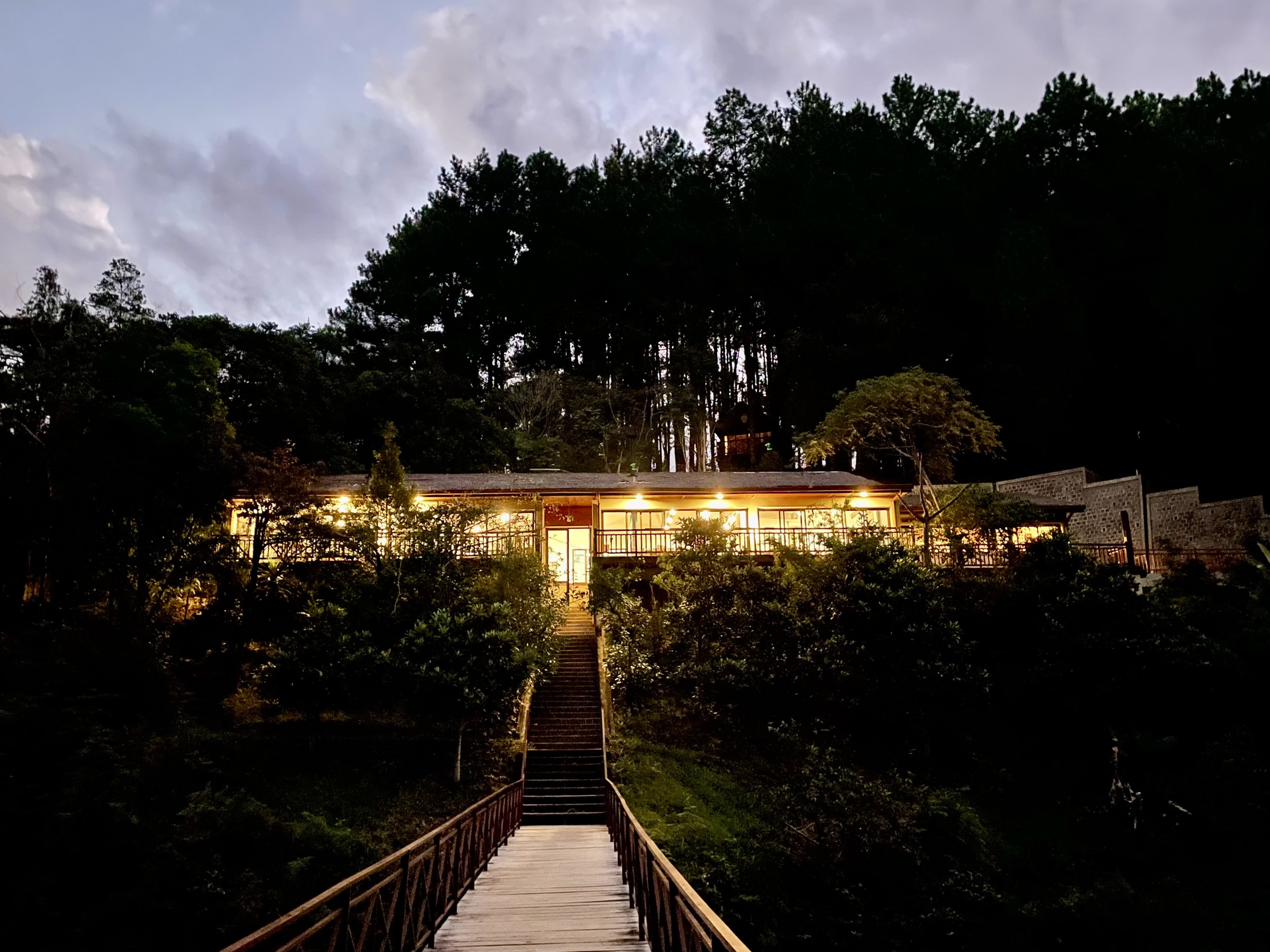
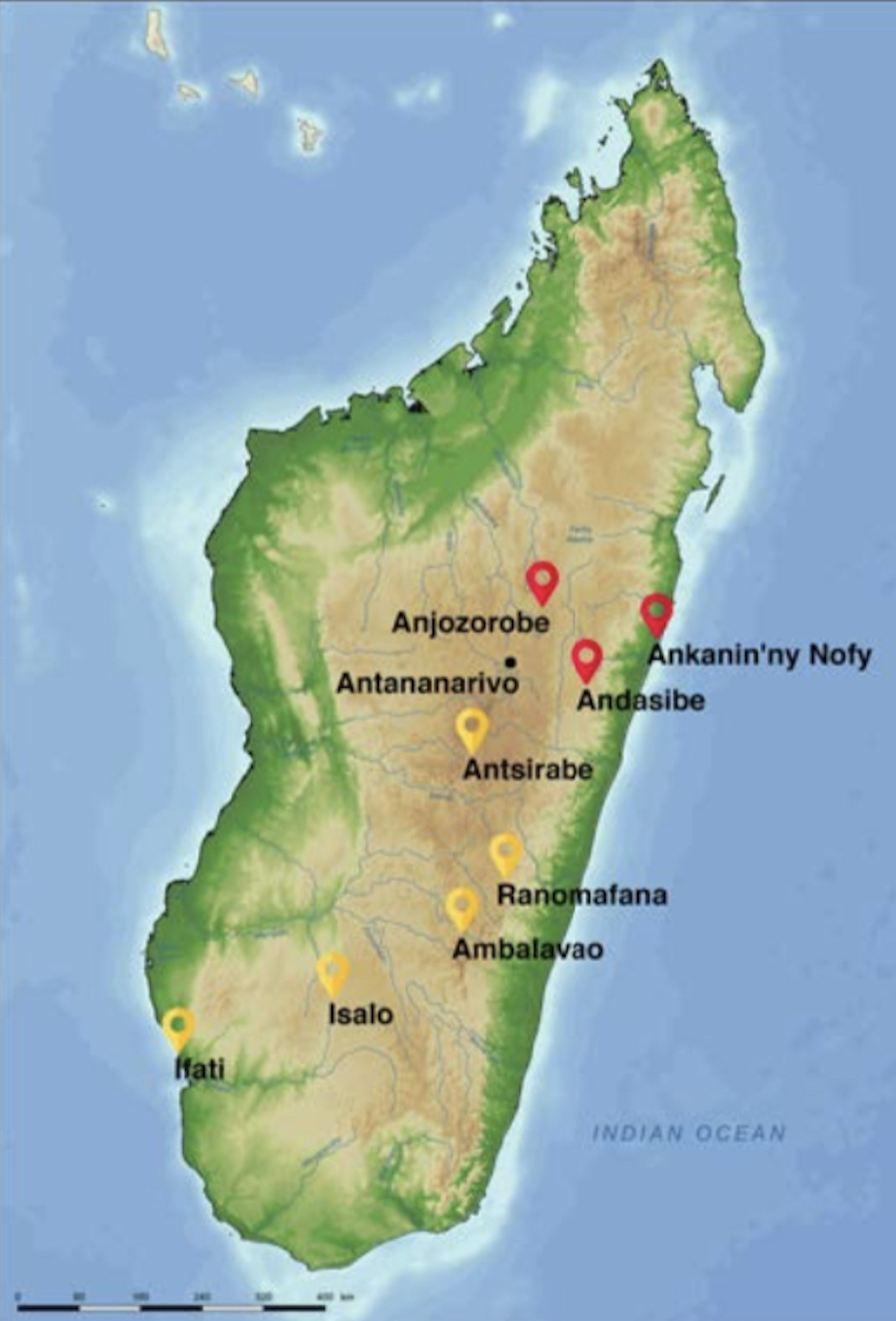
This past April, we embarked on a trip to Madagascar that was 3 years in the making (postponed by COVID-19). Our main three field sites included 1) Anjozorobe, a high plateau forests in Madagascar, 2) Andasibe, which harbors a National Park with lush rainforest, and 3) Ankanin’ny Nofy, a beautiful coastline site located on a peninsula in eastern Madagascar. Below are photos highlighting some of our incredible wildlife sightings, from chameleons, to lemurs and of course so many unique insects and other amazing invertebrates.
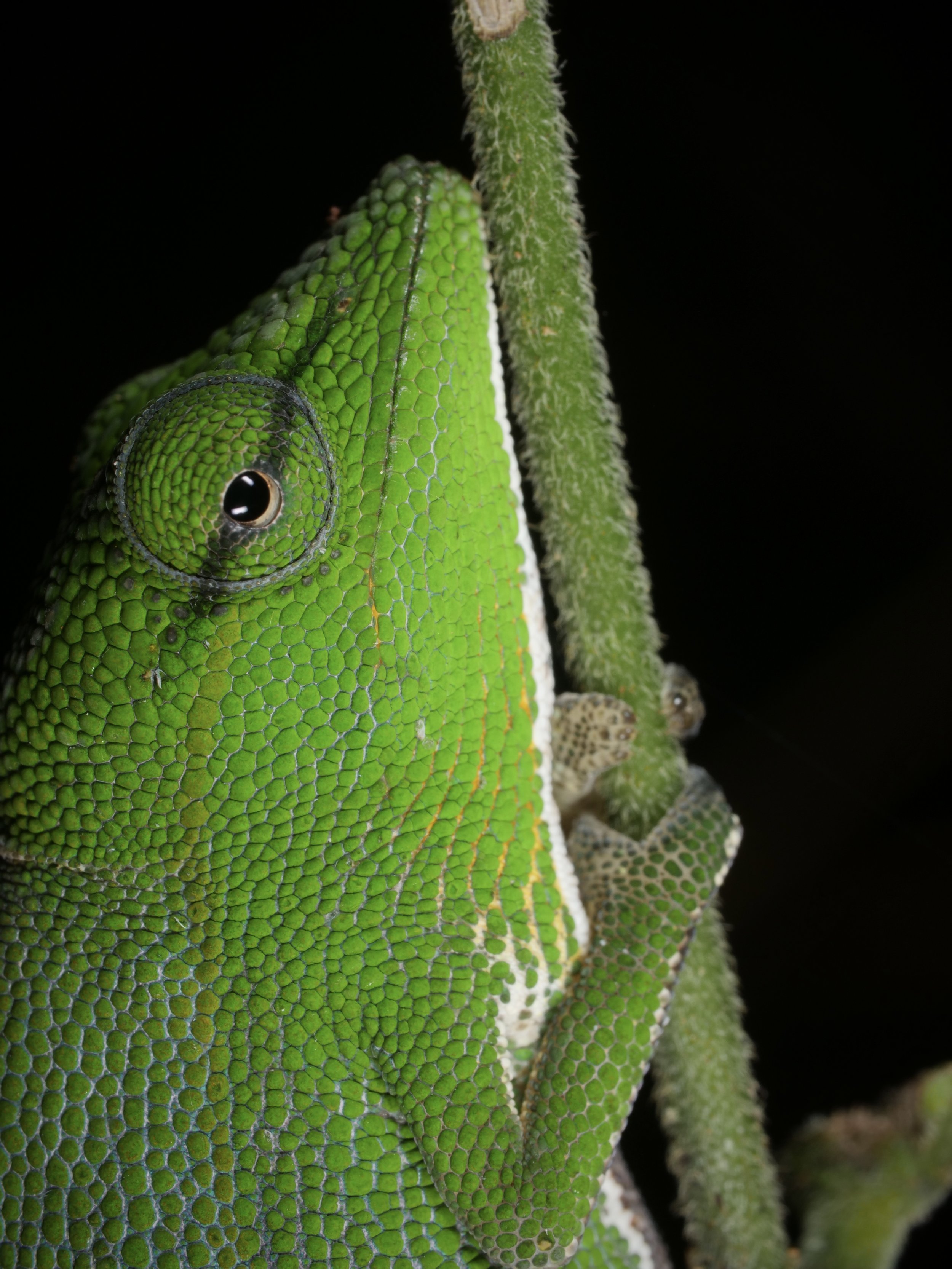
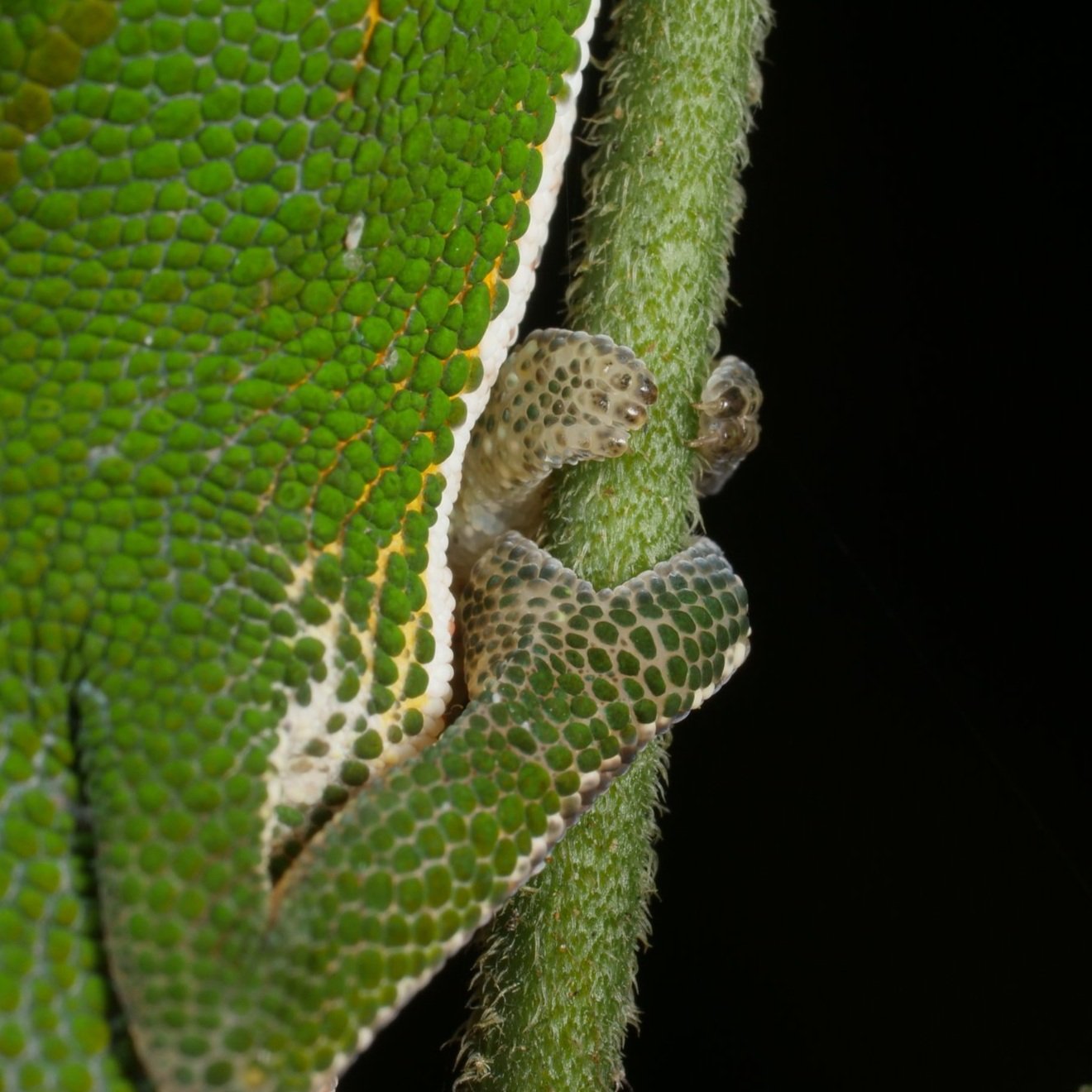


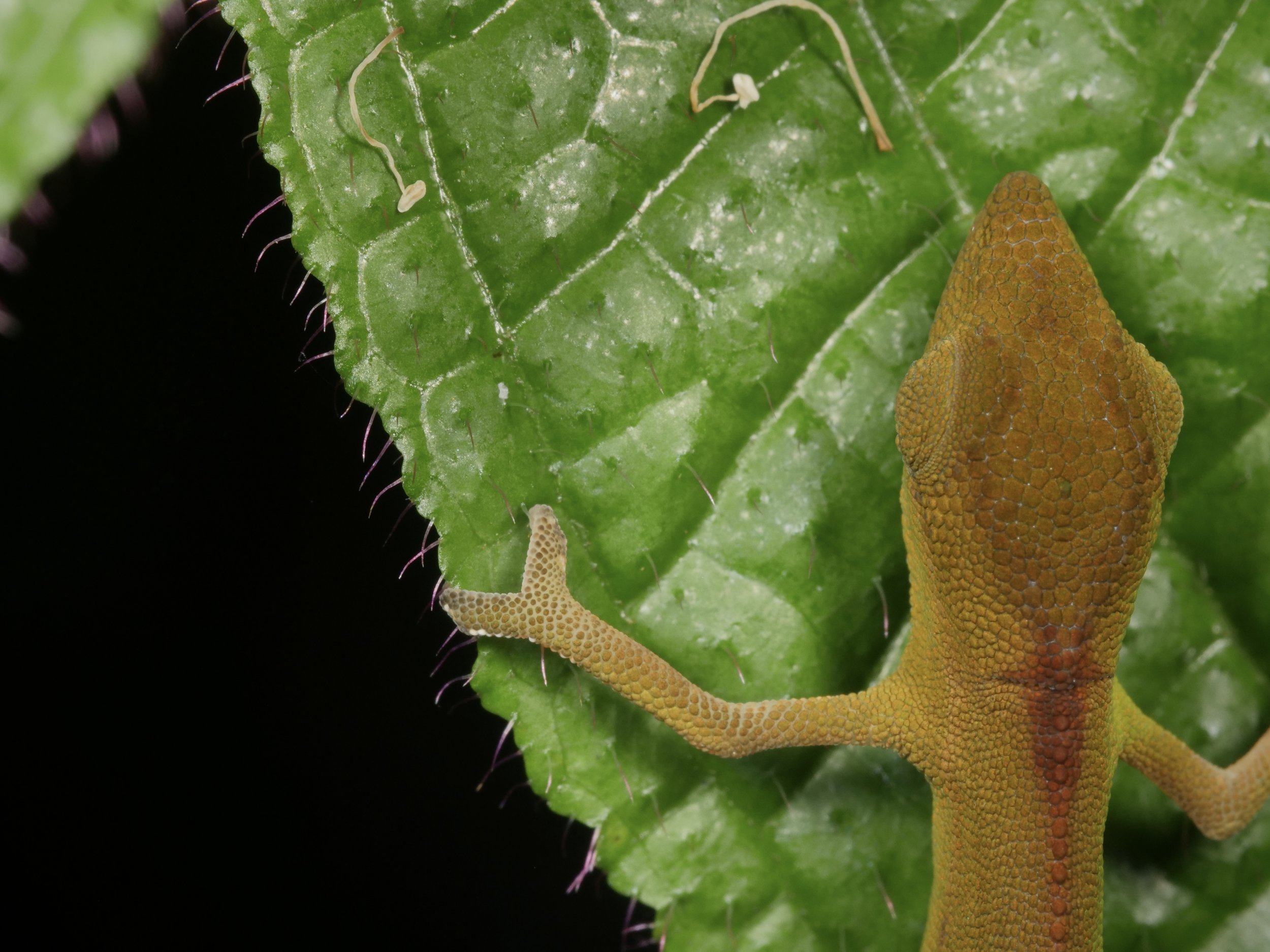
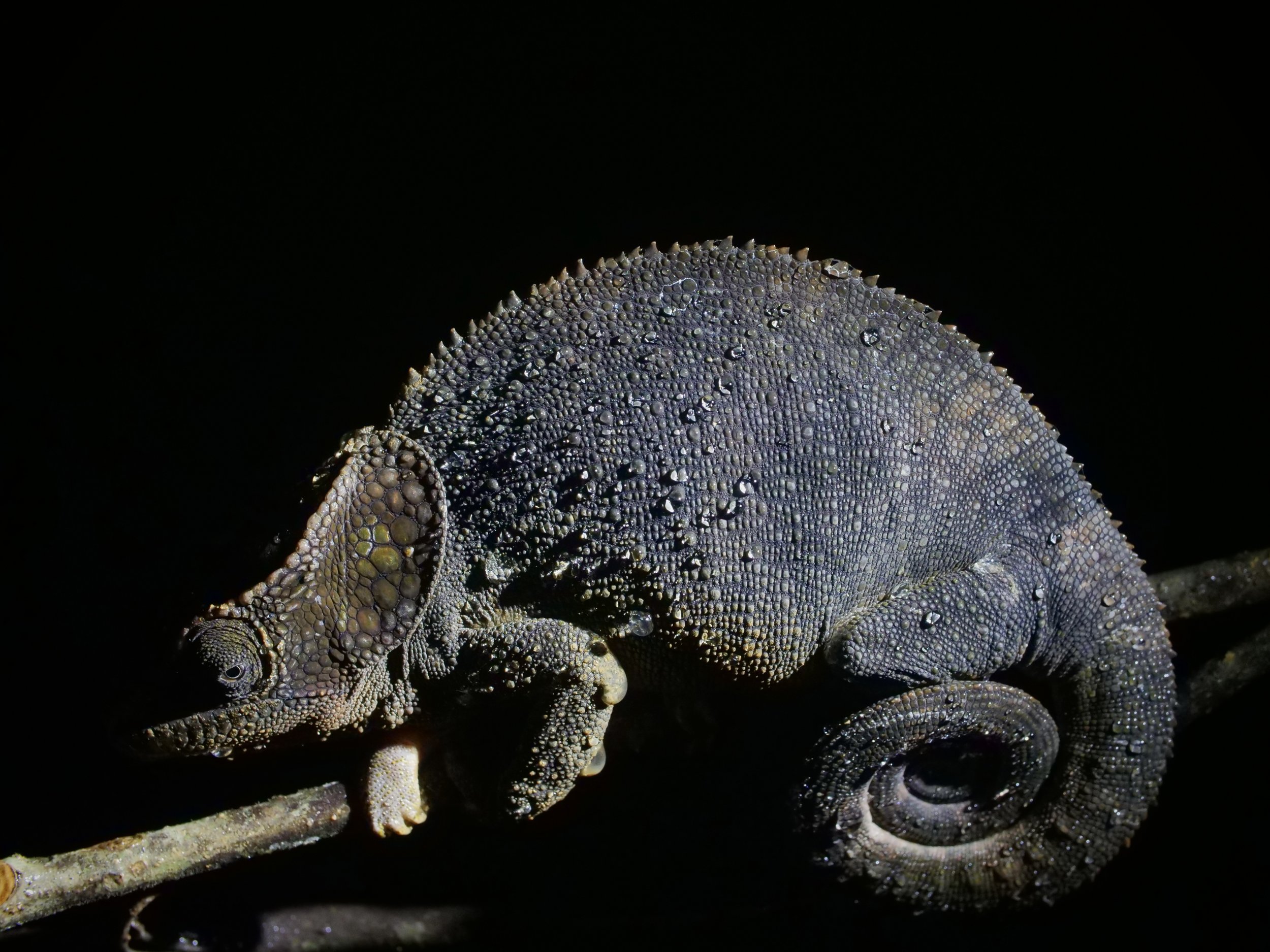
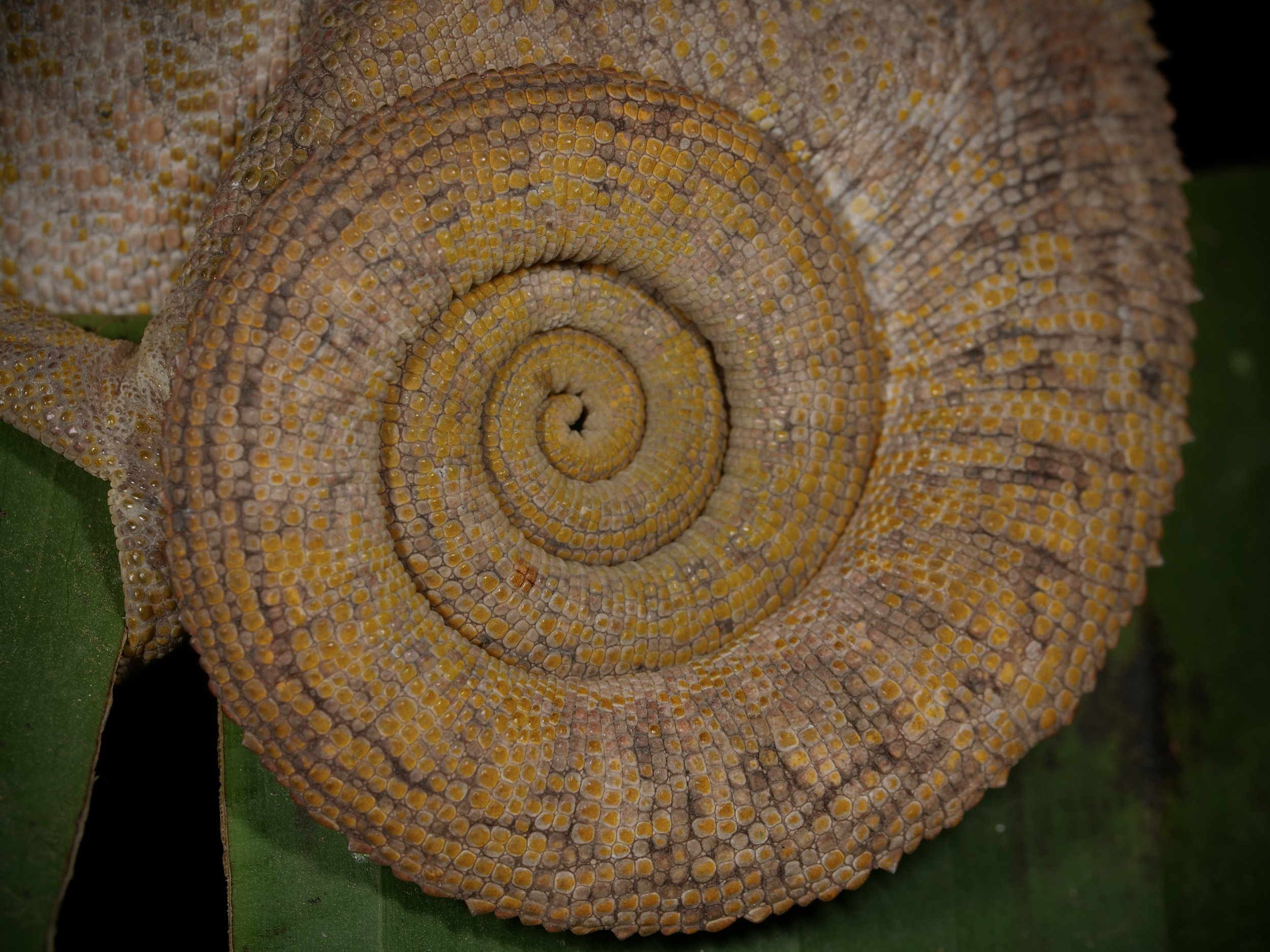
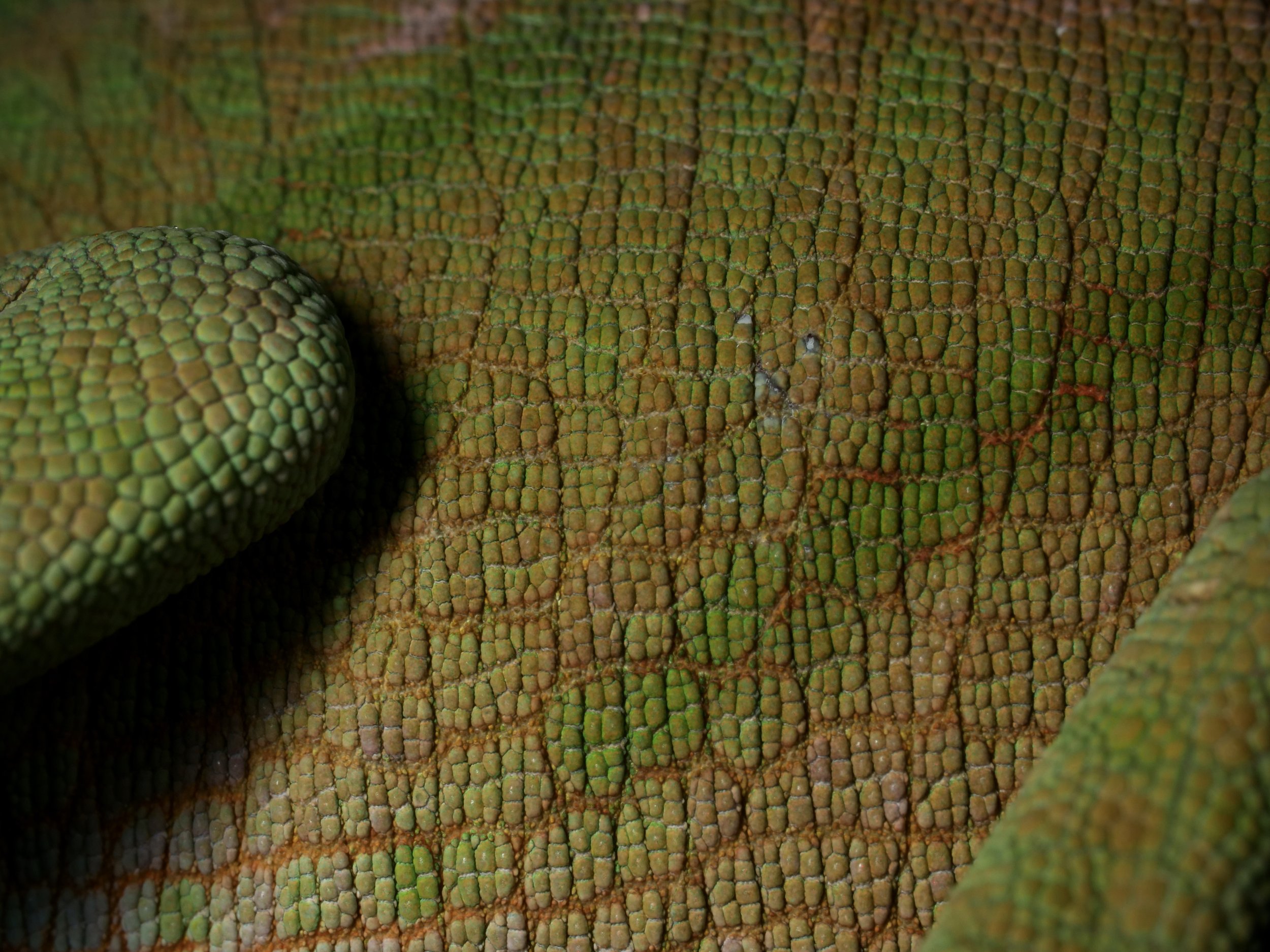
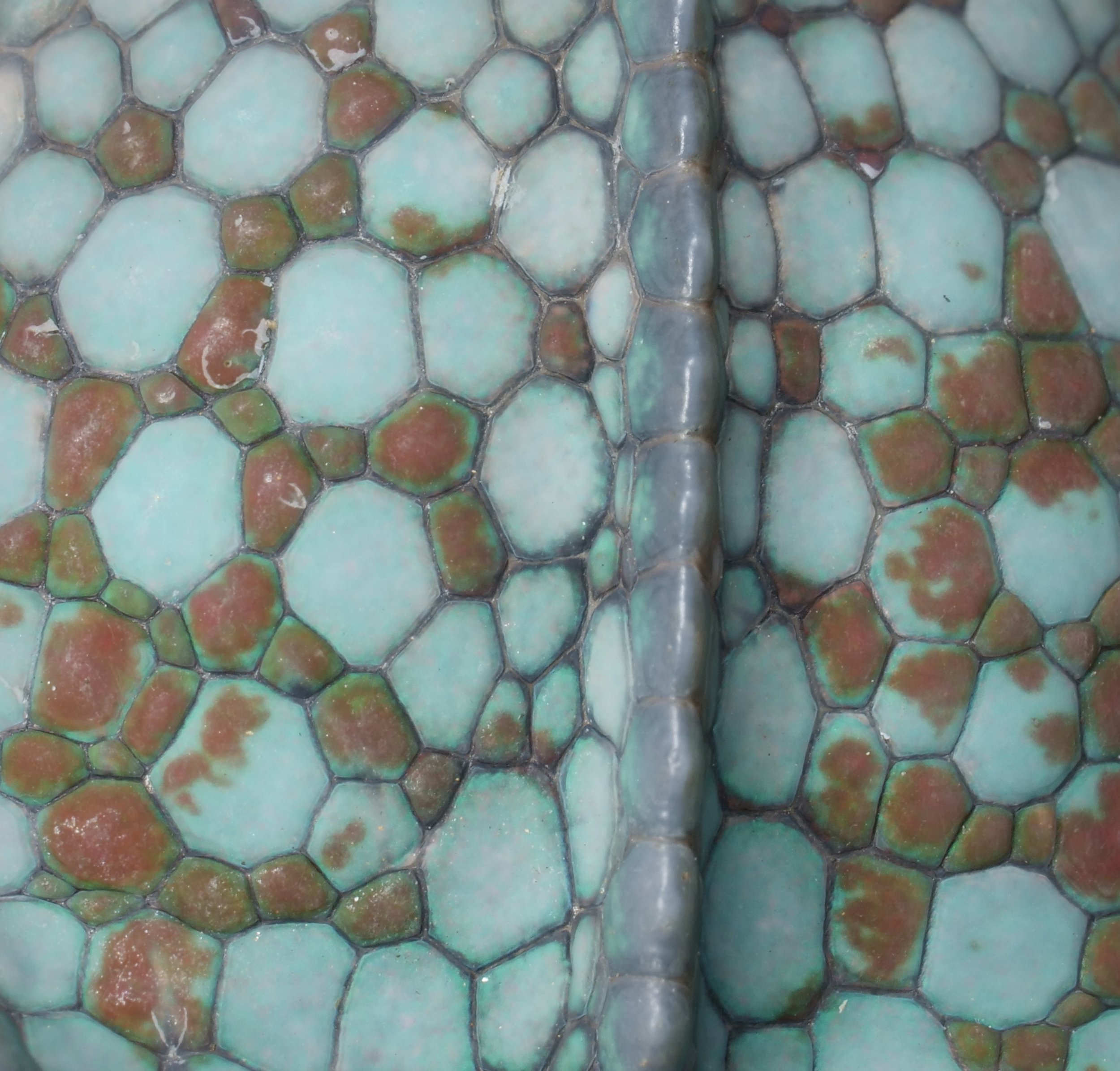
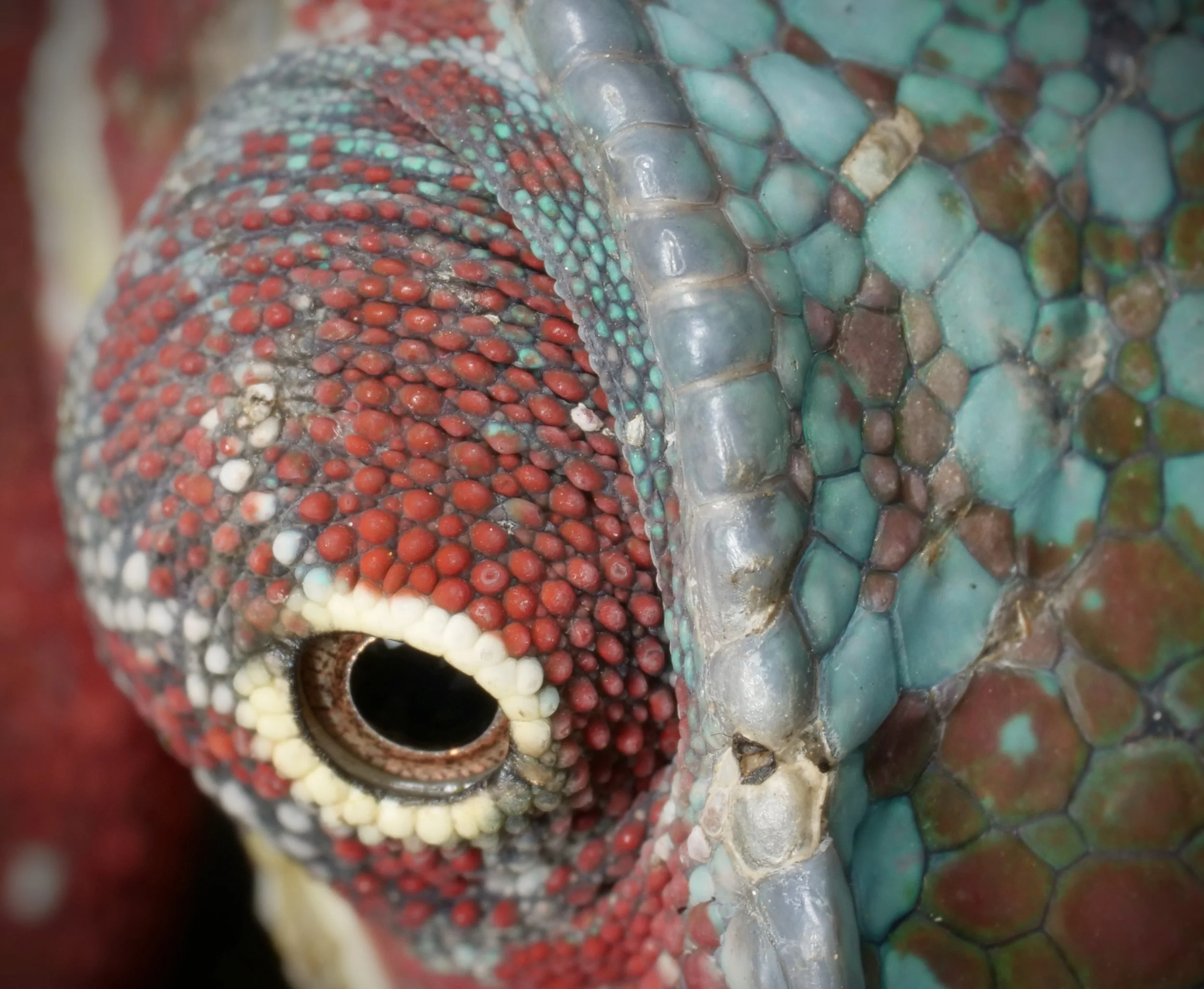
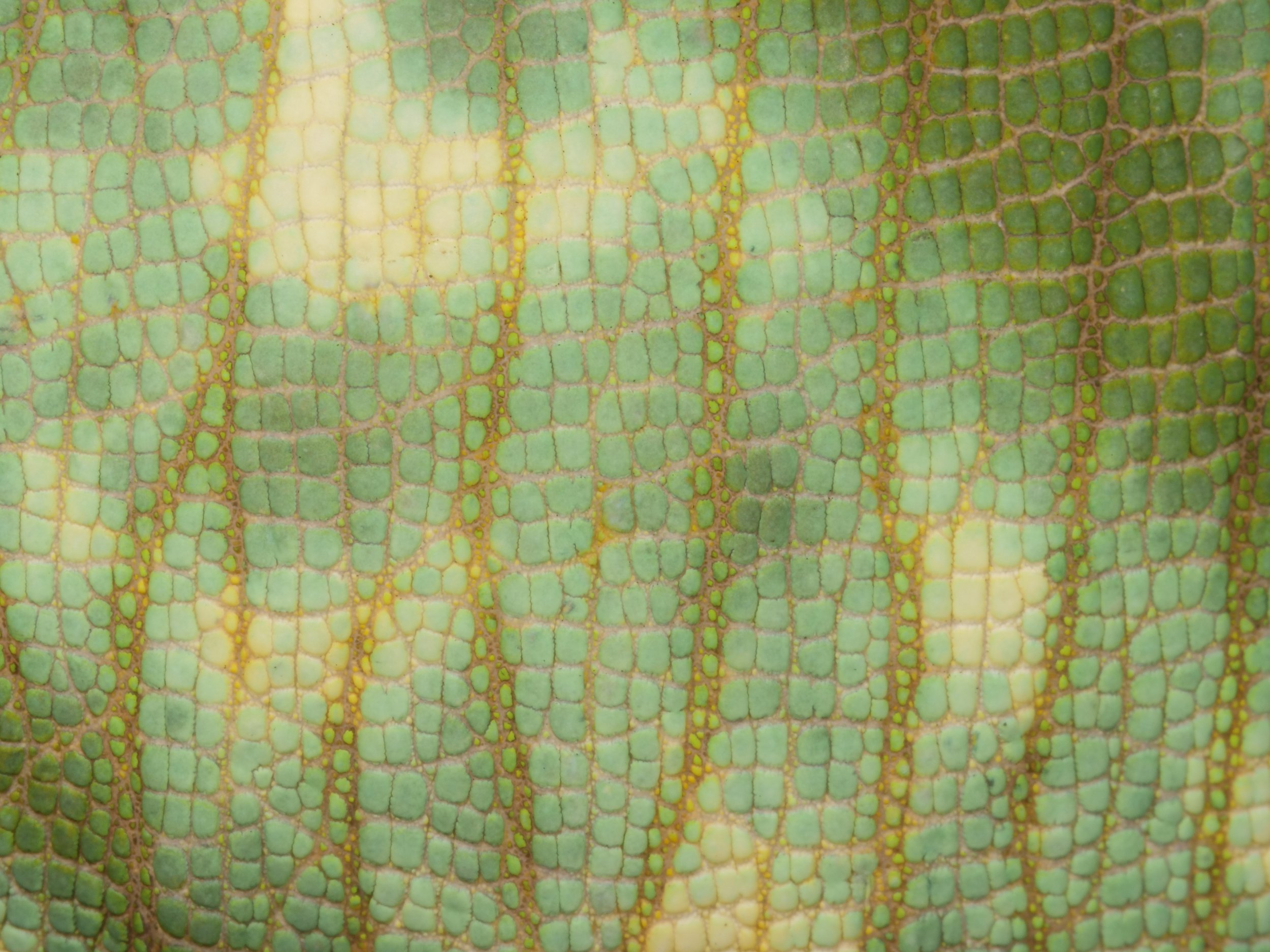
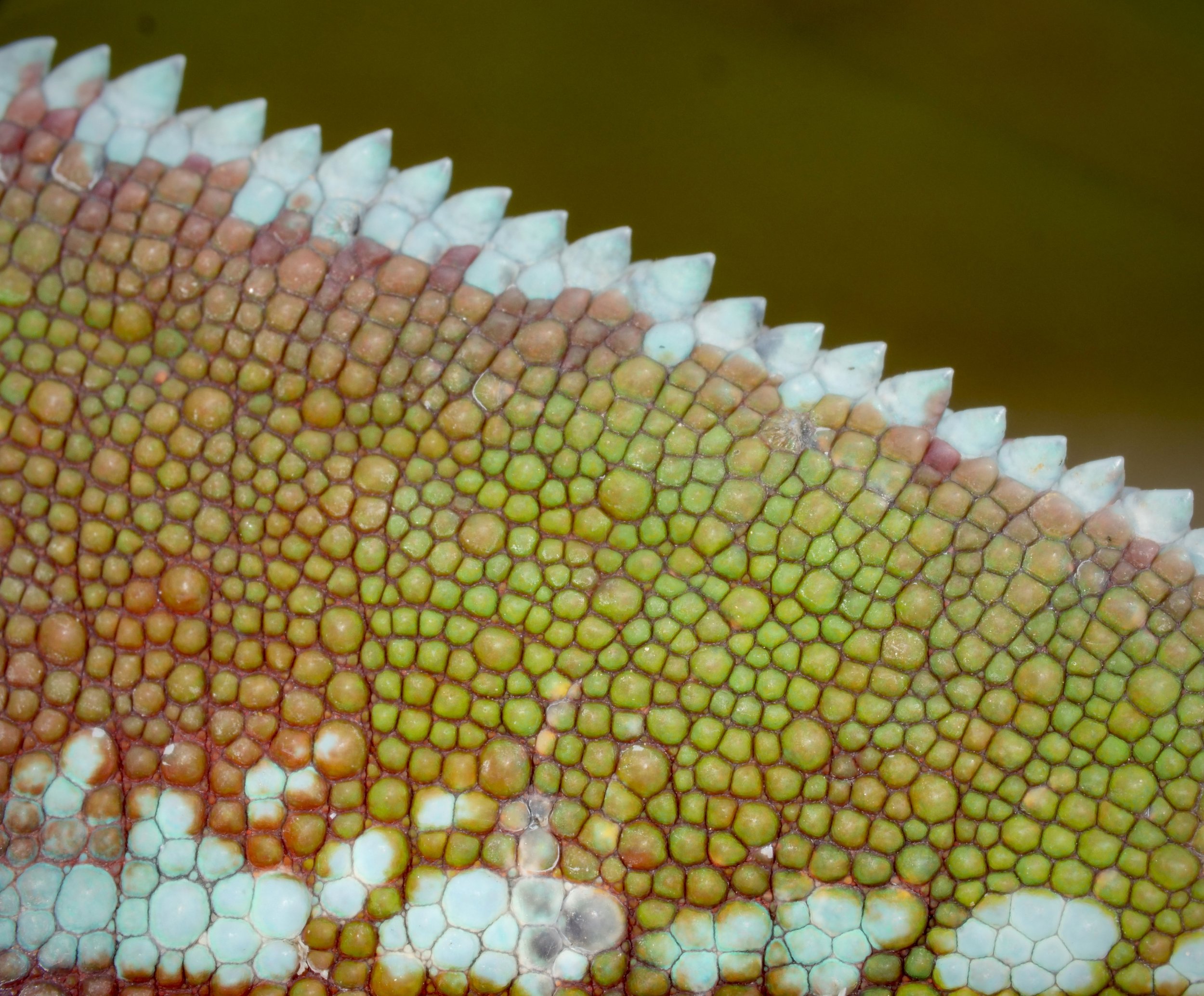
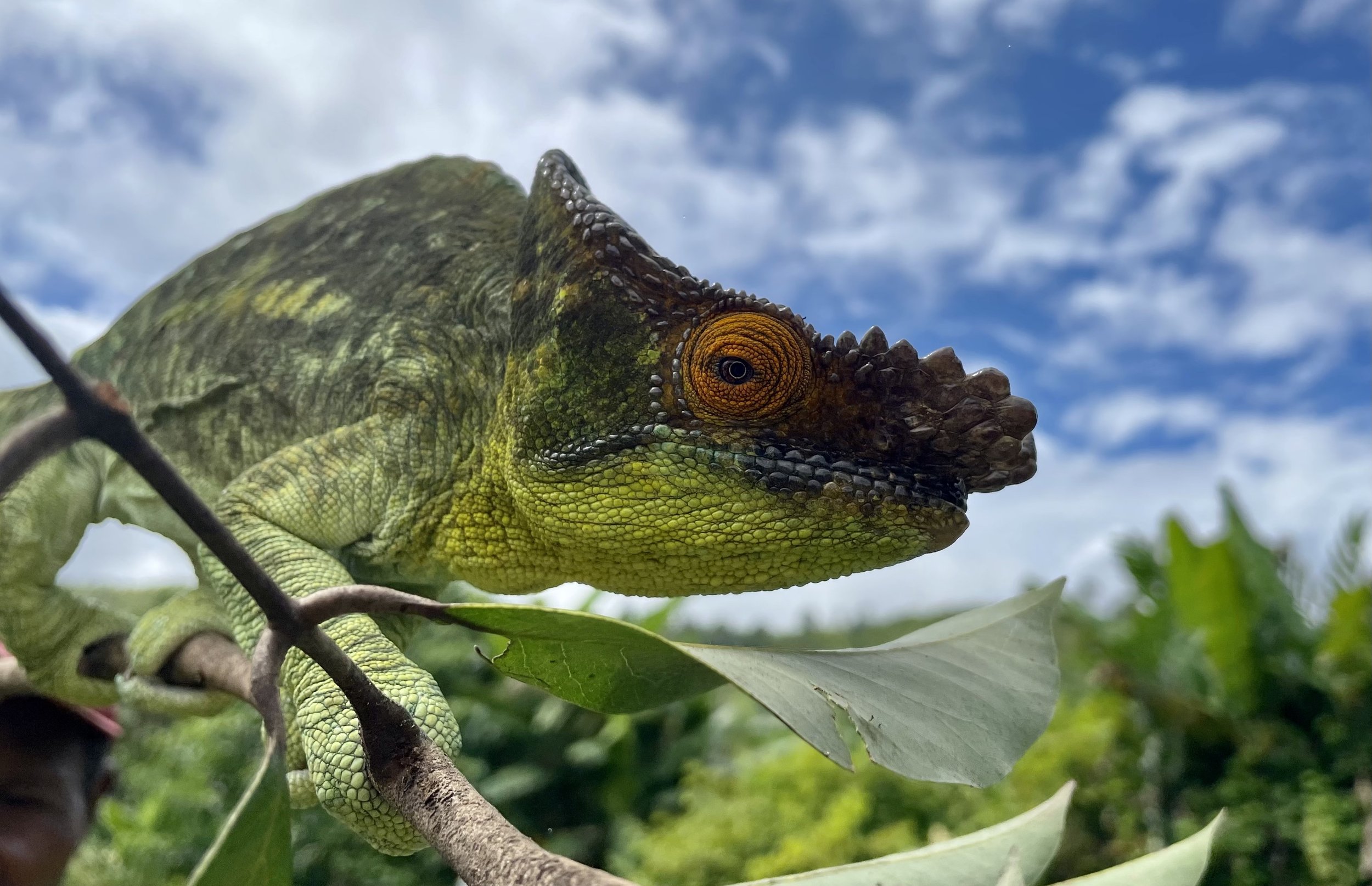
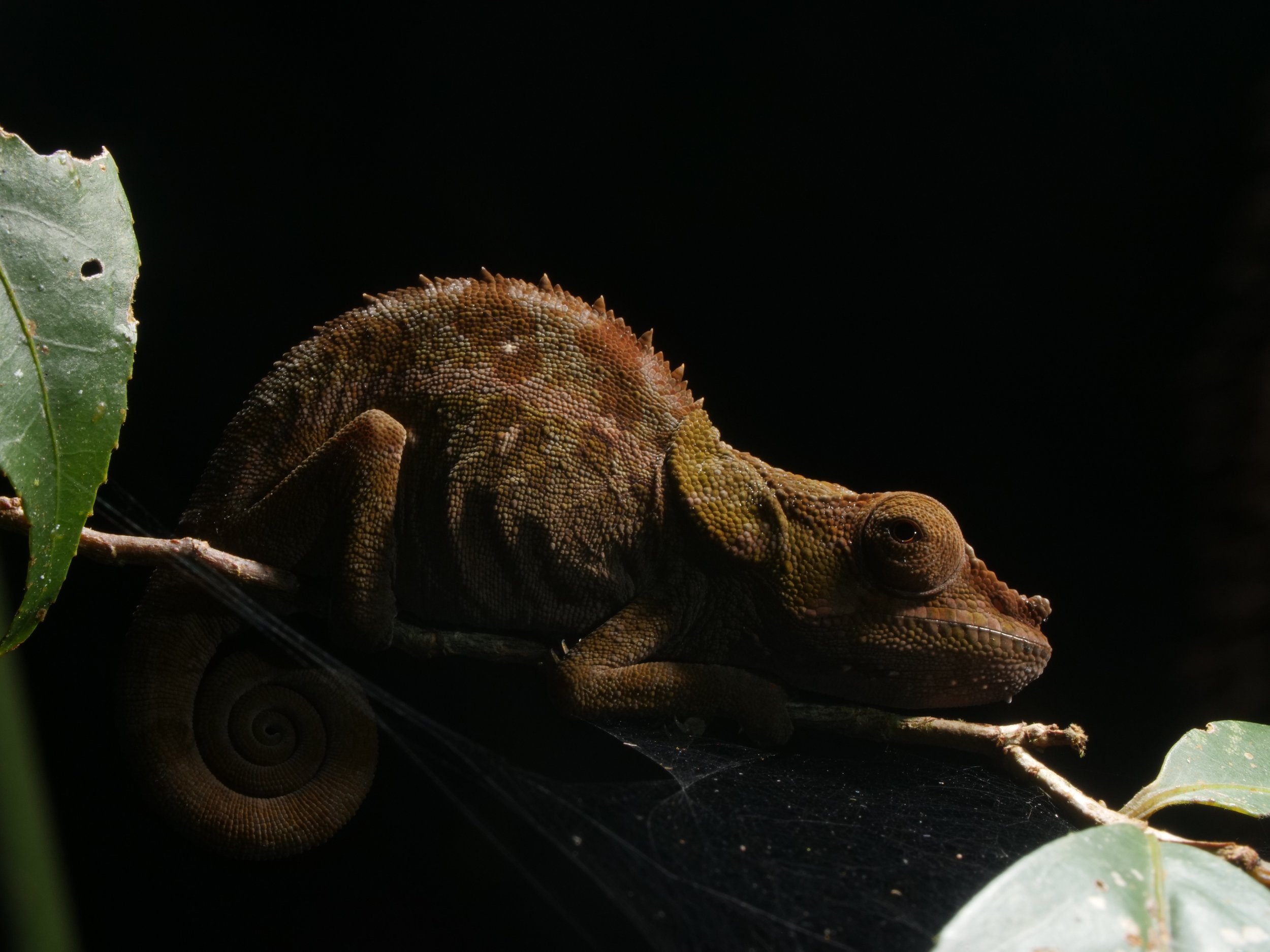
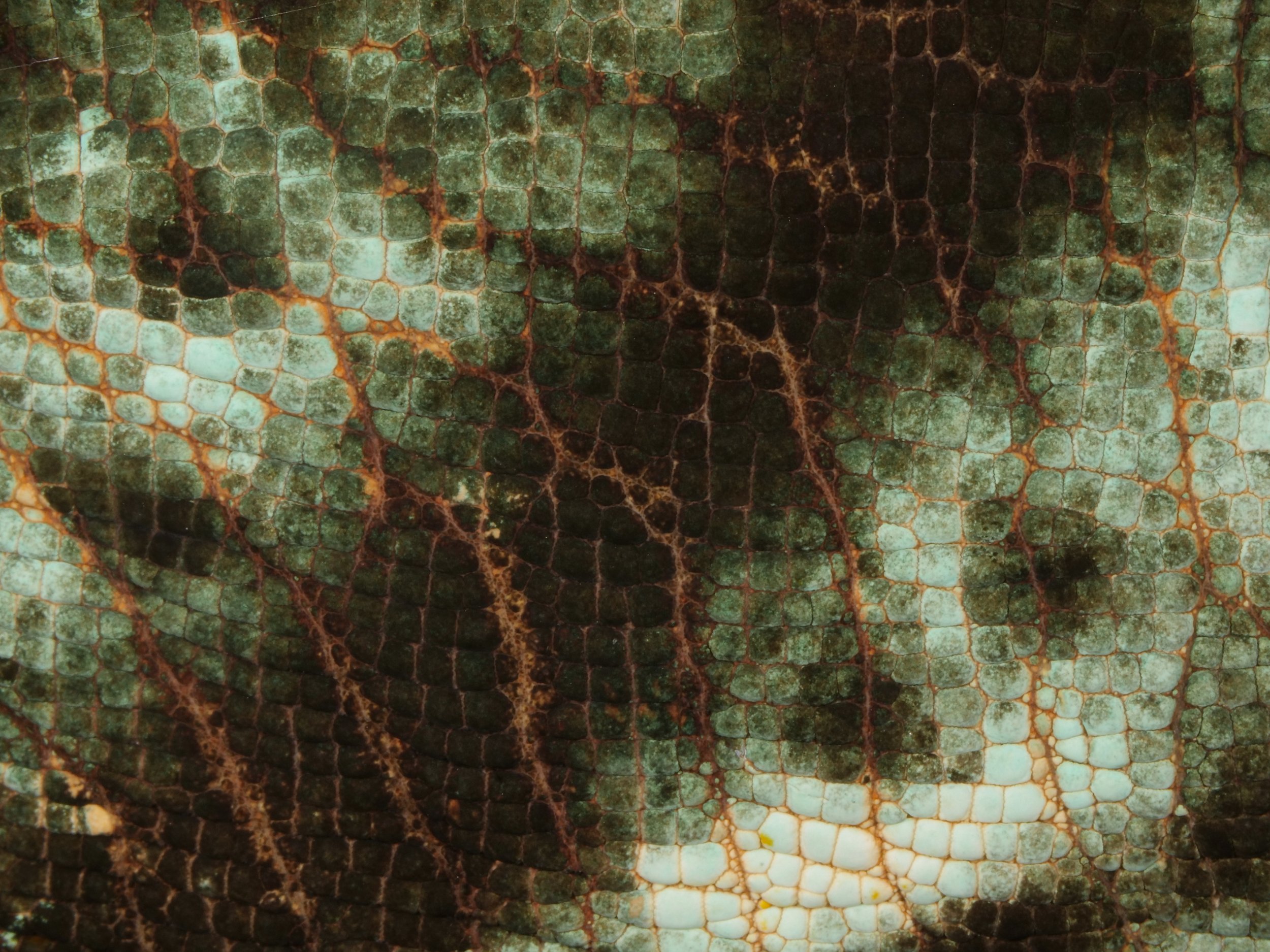
Zooming in on the scales of a Perinet chameleon, also known as the Malagasy side-striped chameleon.
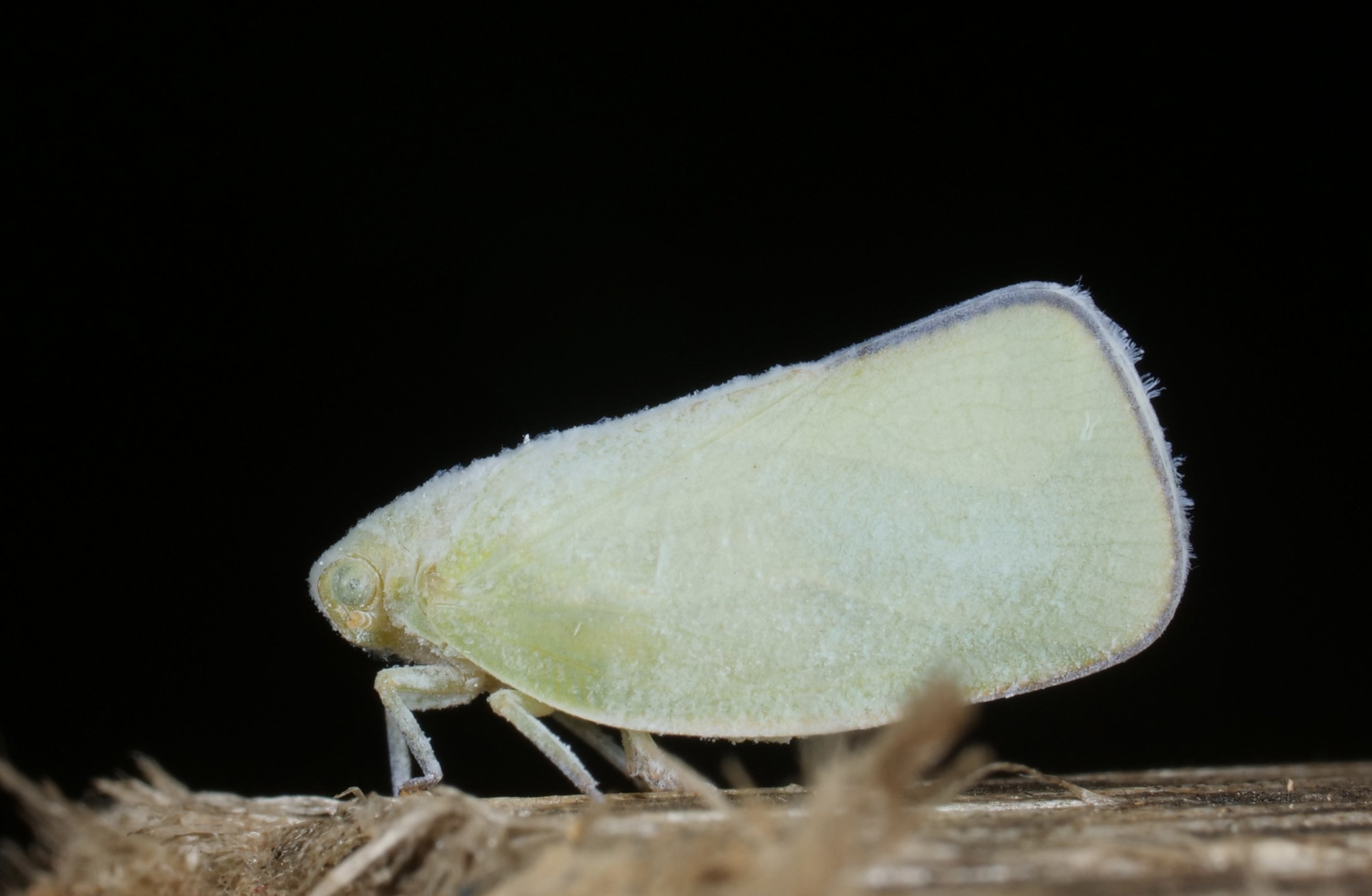
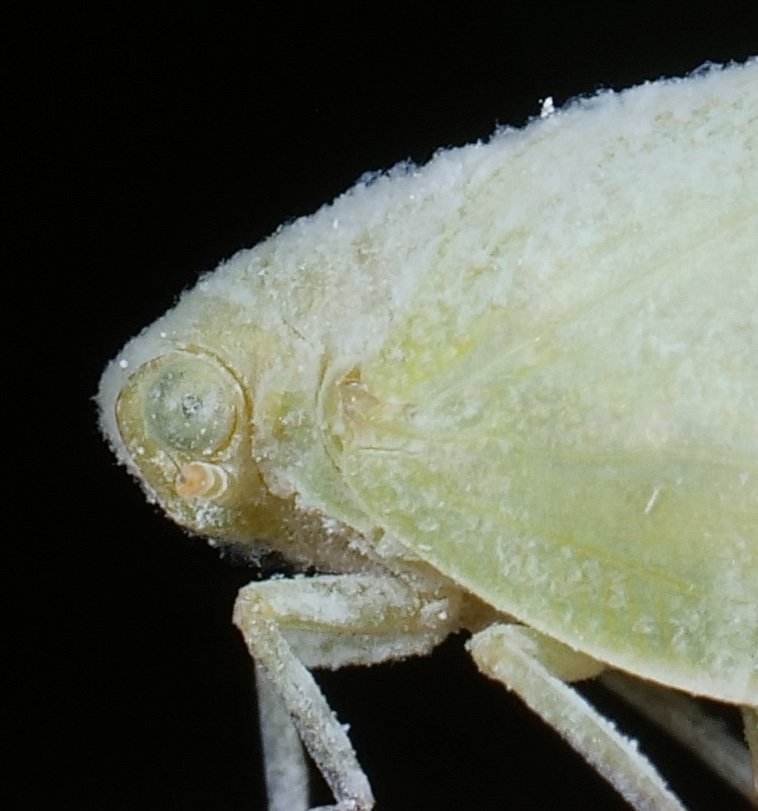
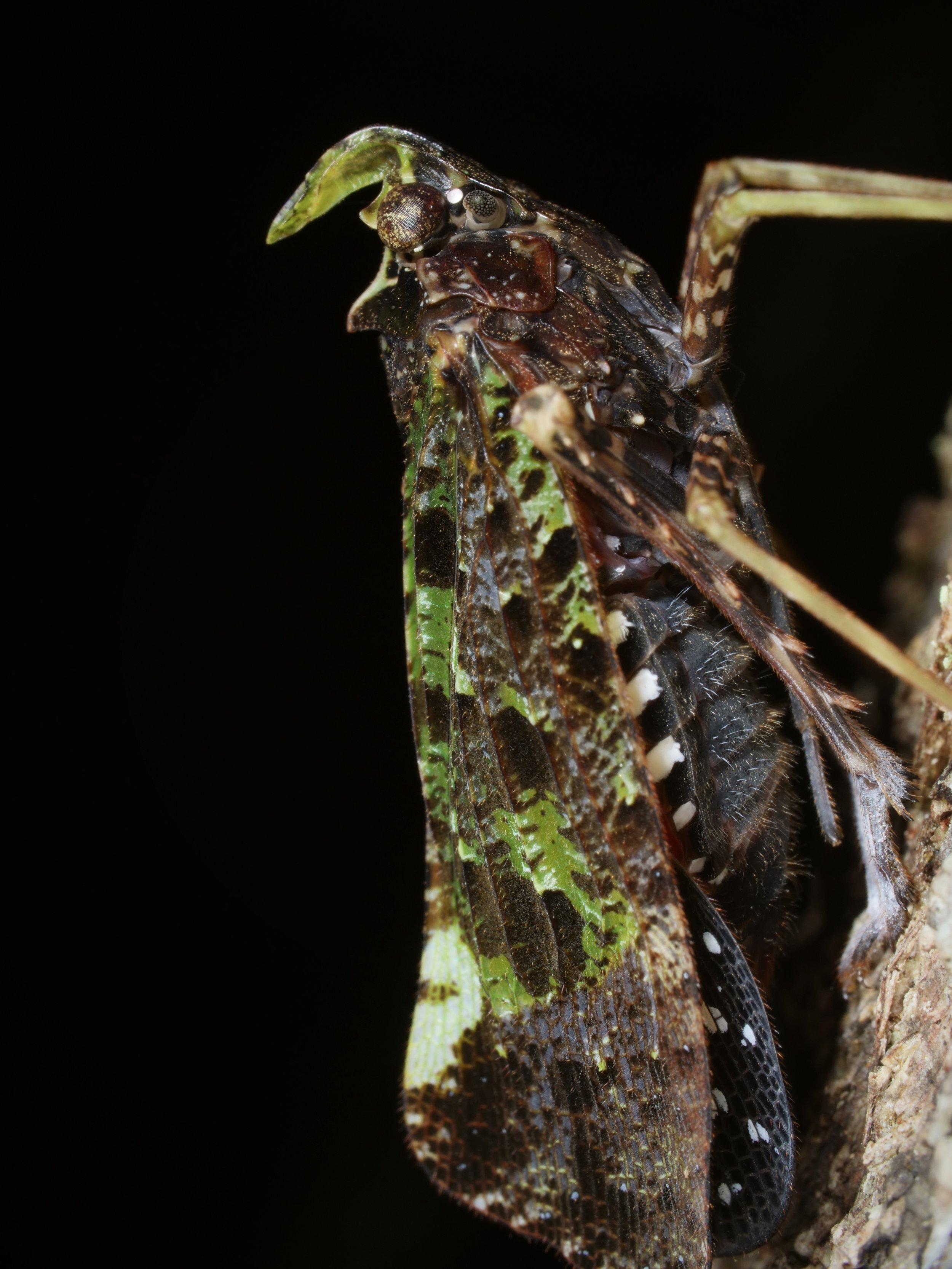
A waxy planthopper
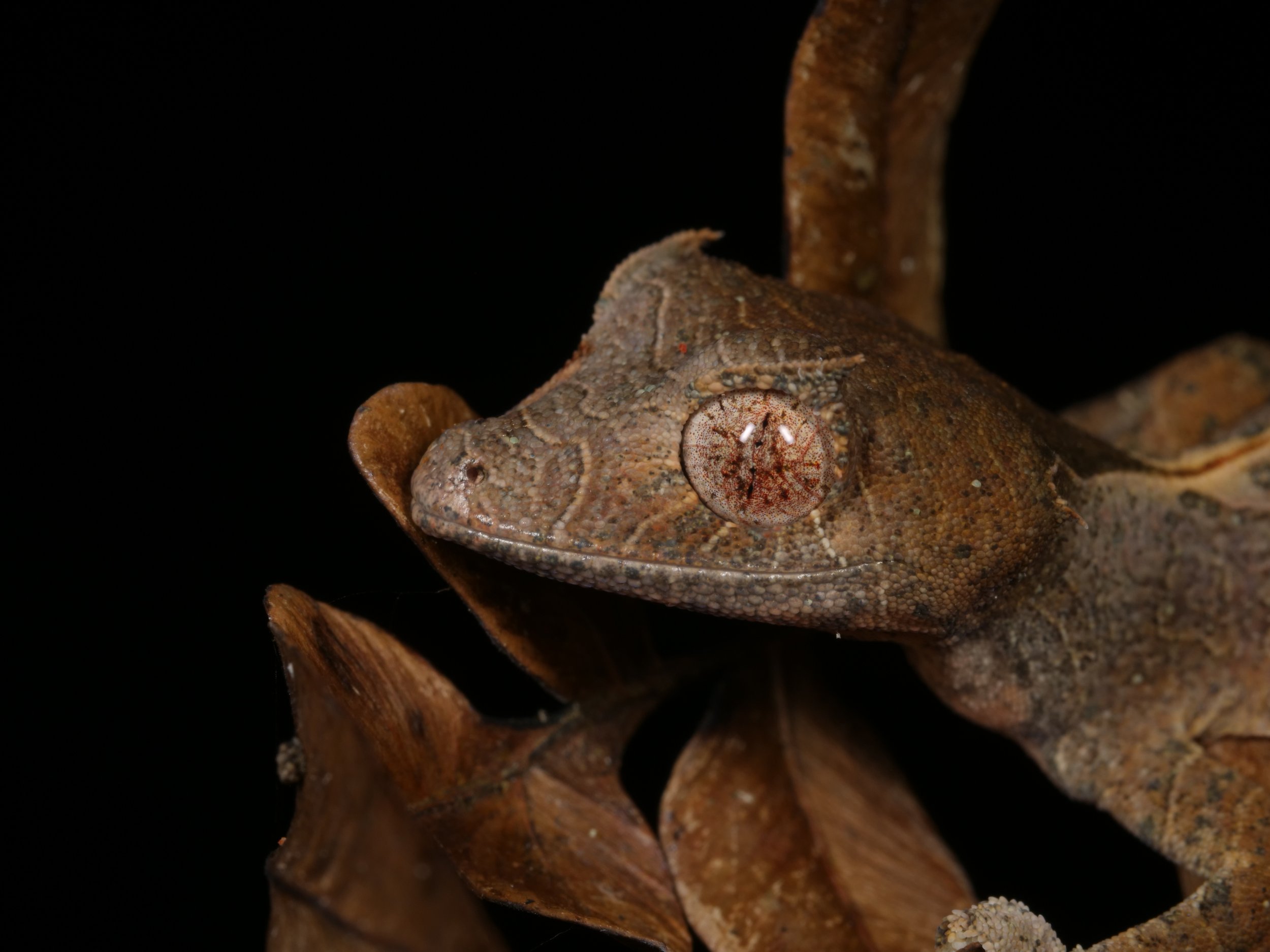
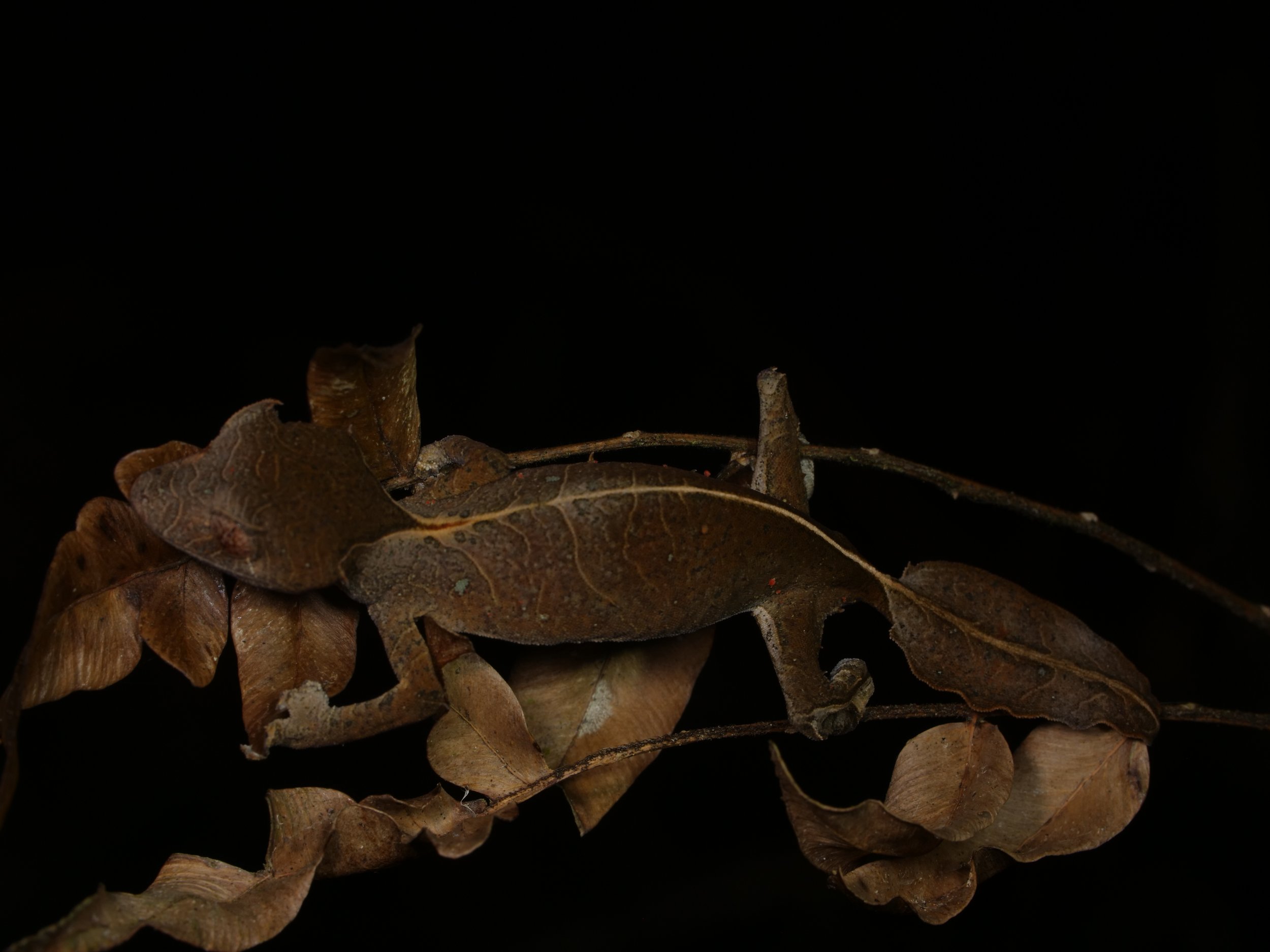
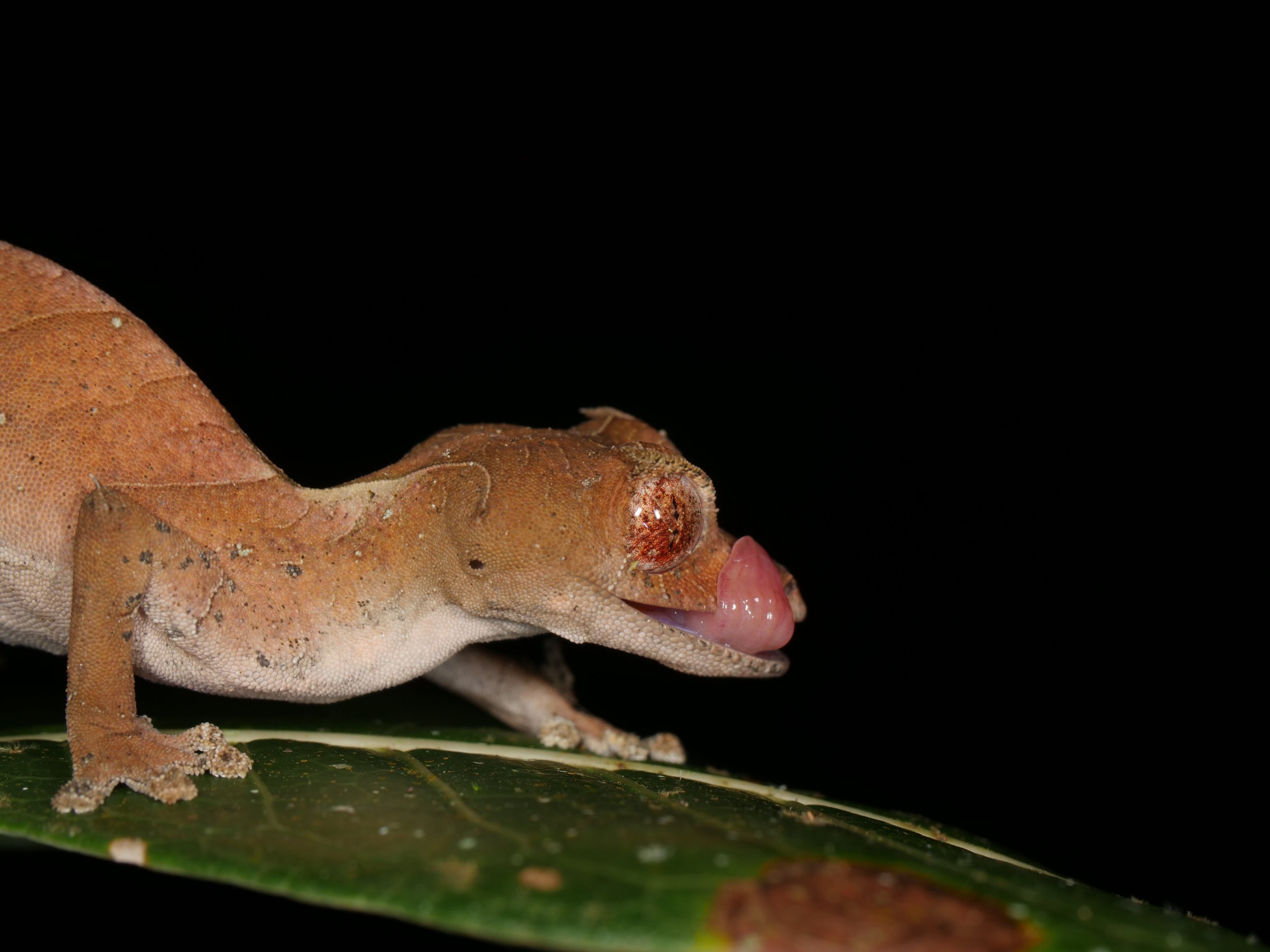
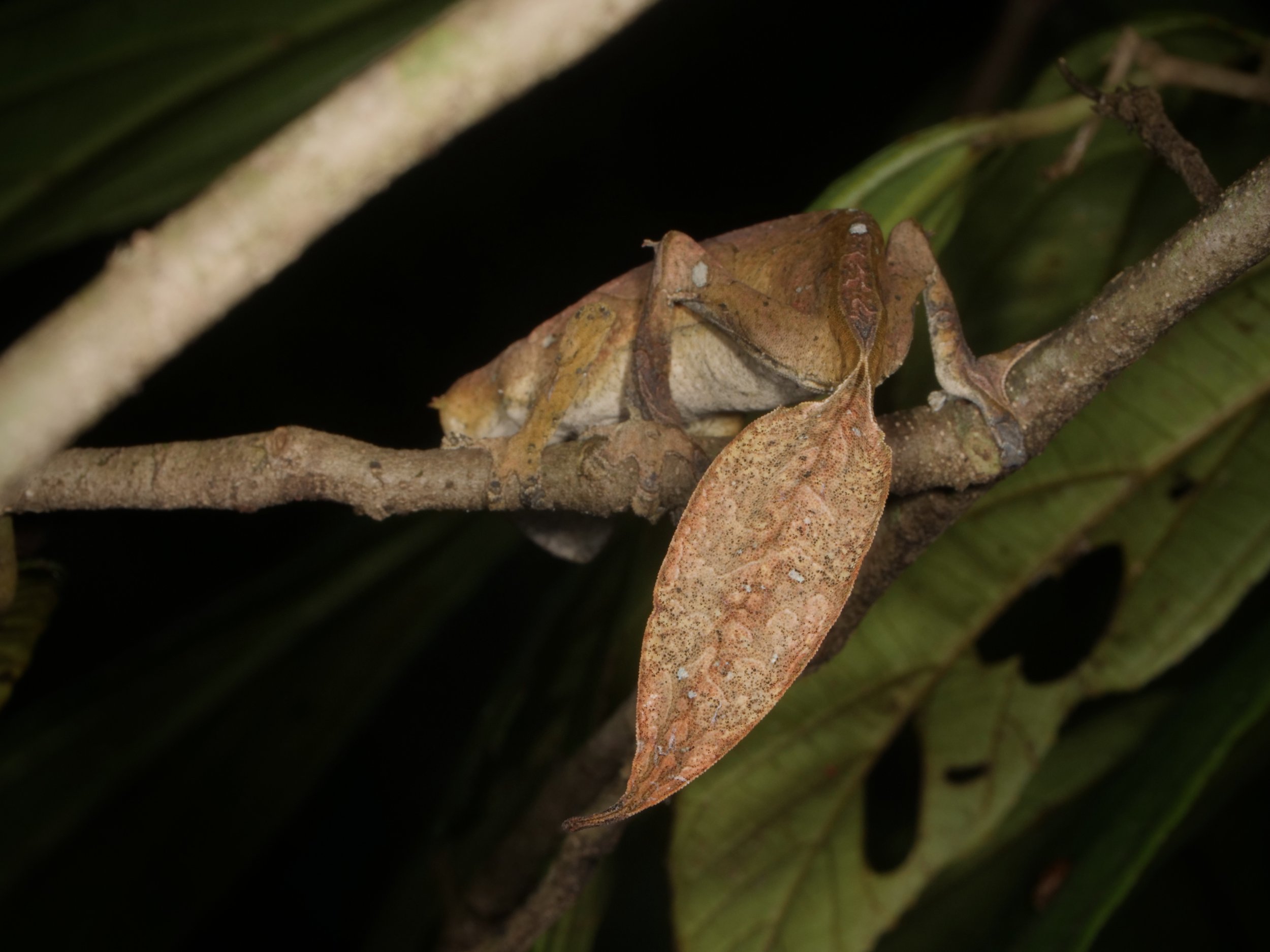
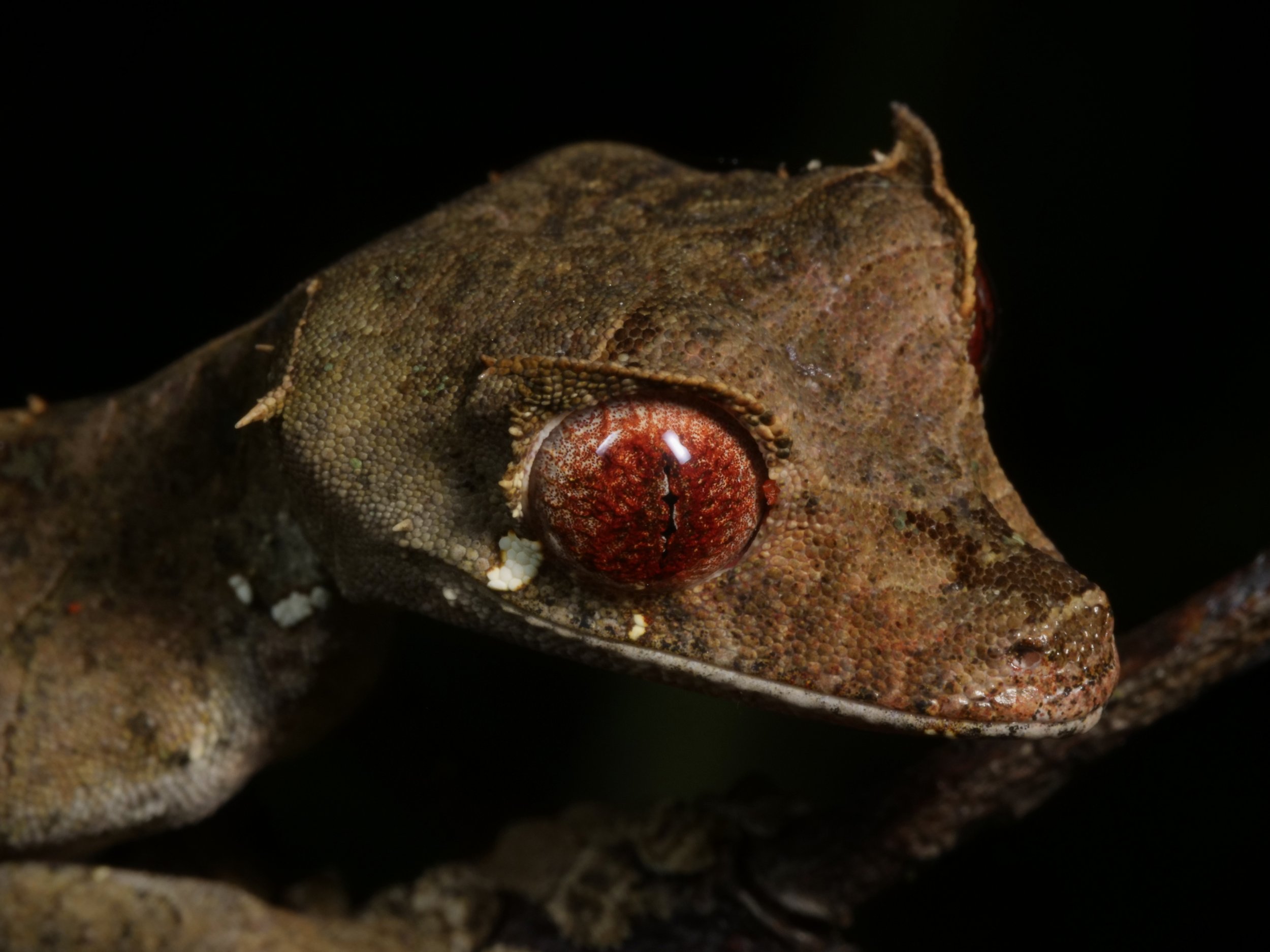





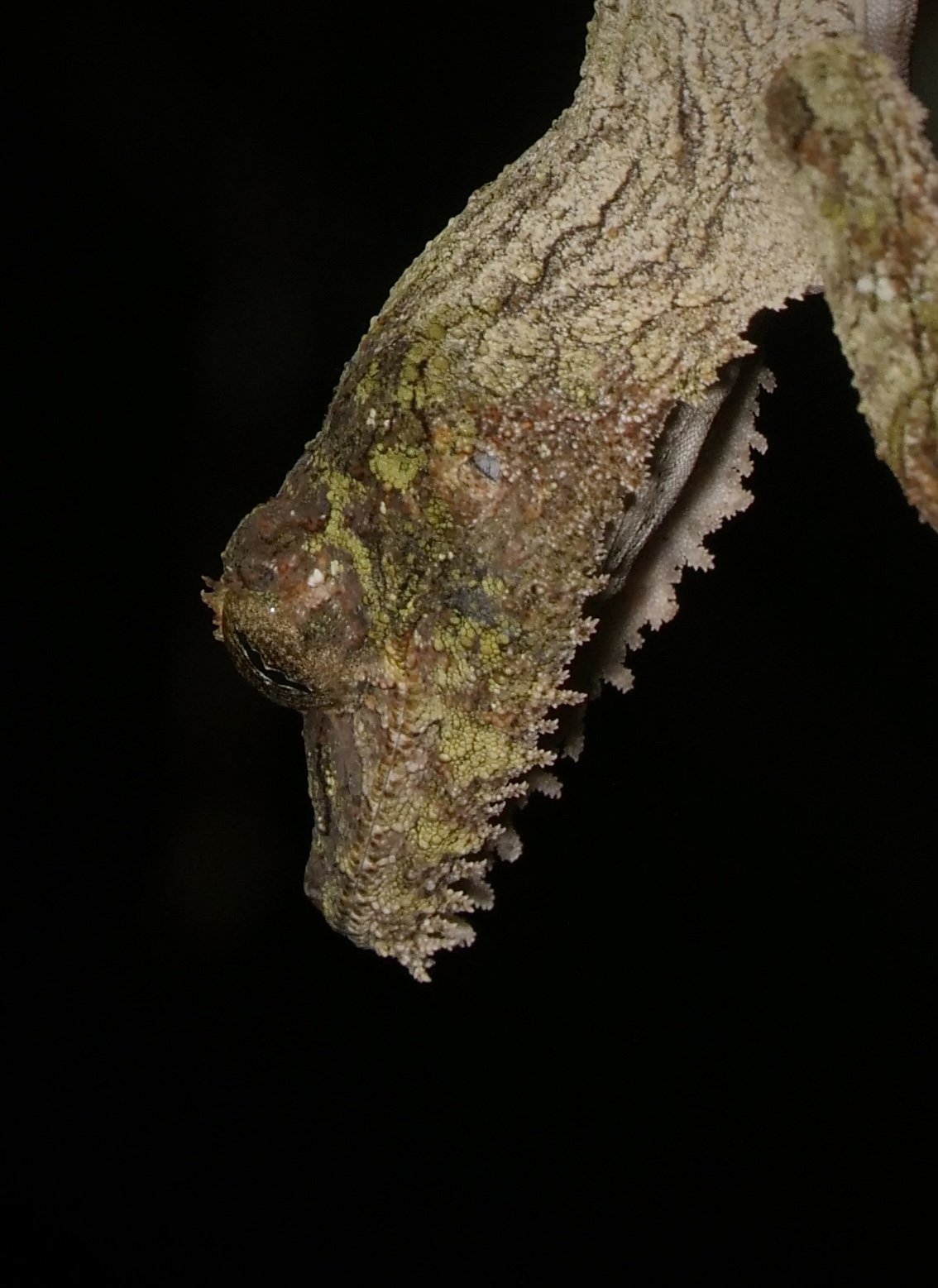
We spotted several of these wonderful satanic leaf-tailed geckos, Uroplatus phantasticus, a species indigenous to the island of Madagascar. As the name implies, their tails are spitting images of flat, dead leaves, down to the tiniest of details for camouflage.
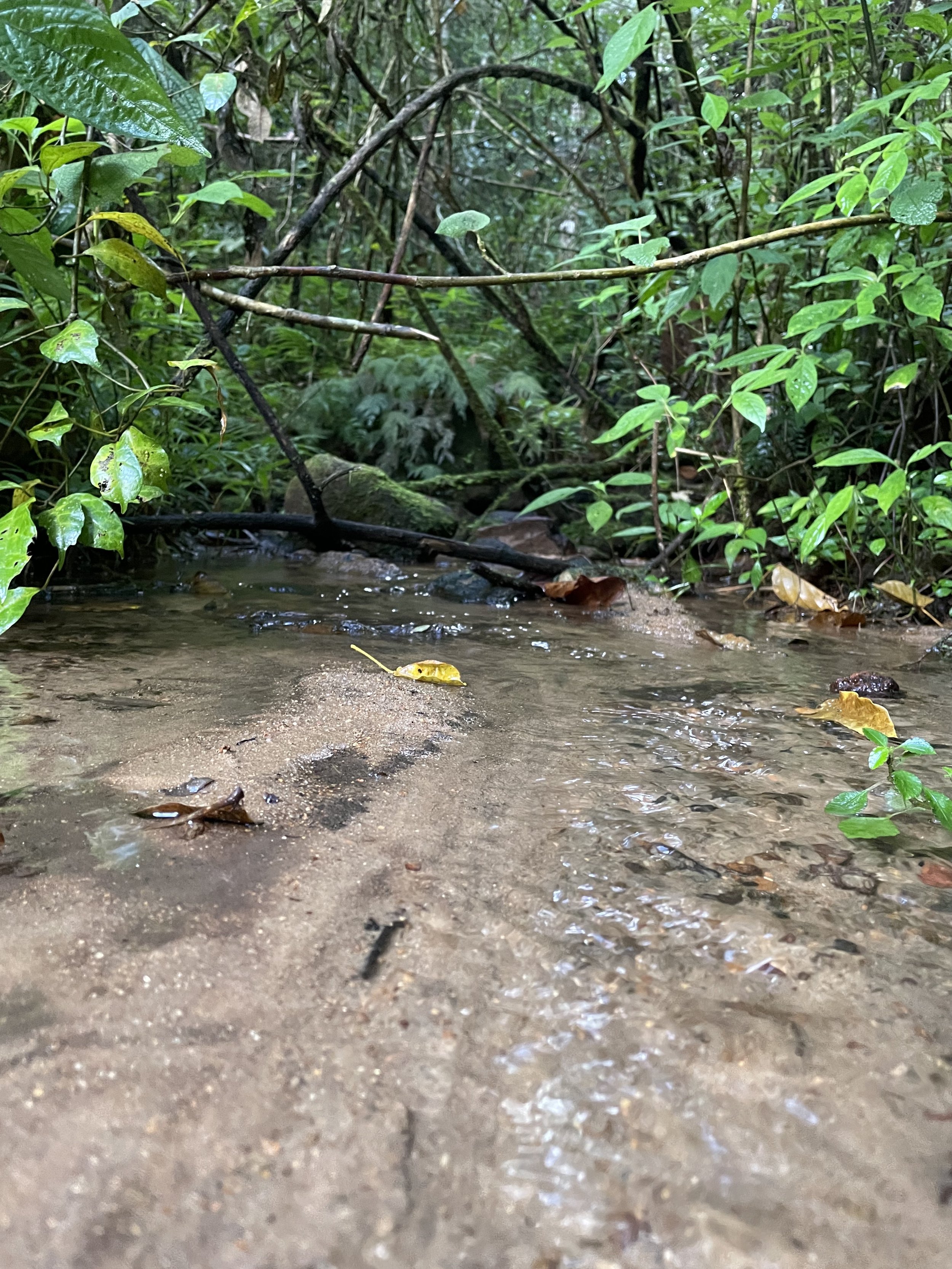
Following our stay in Anjozorobe, we next set out to the lush rainforests of Andasibe. In this locality, up to 100 species of amphibians and 30 species of snakes have been recorded.
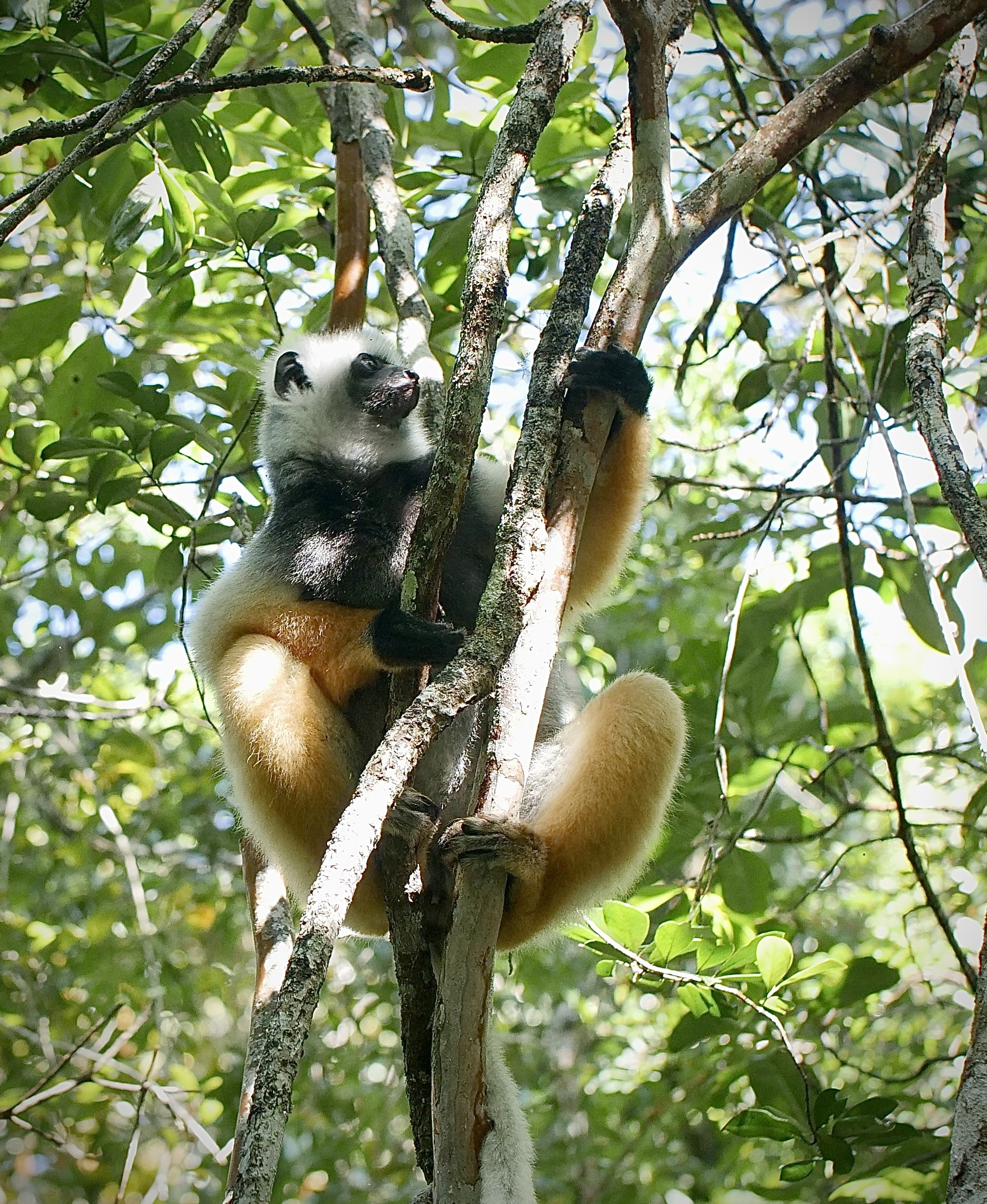
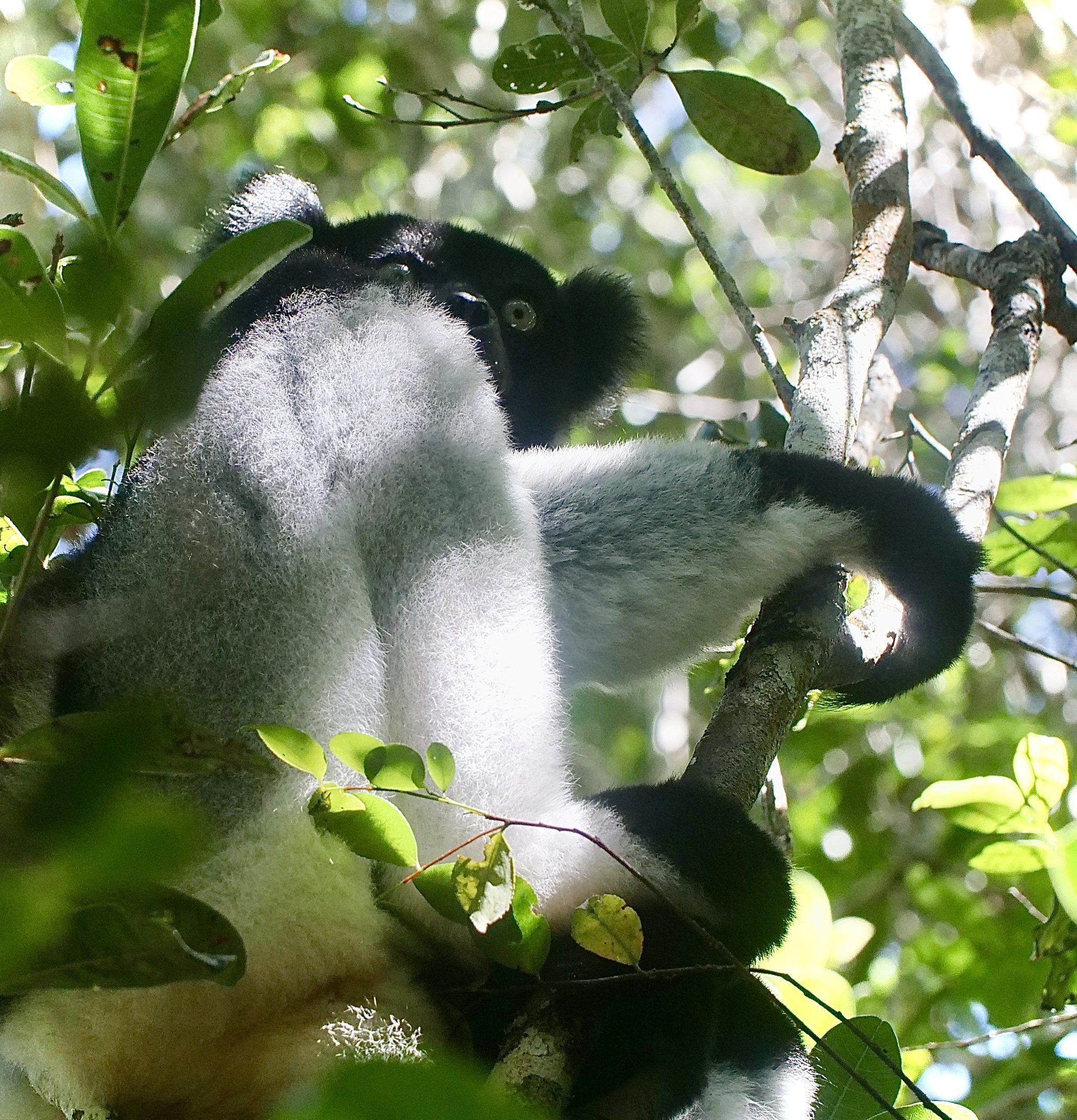
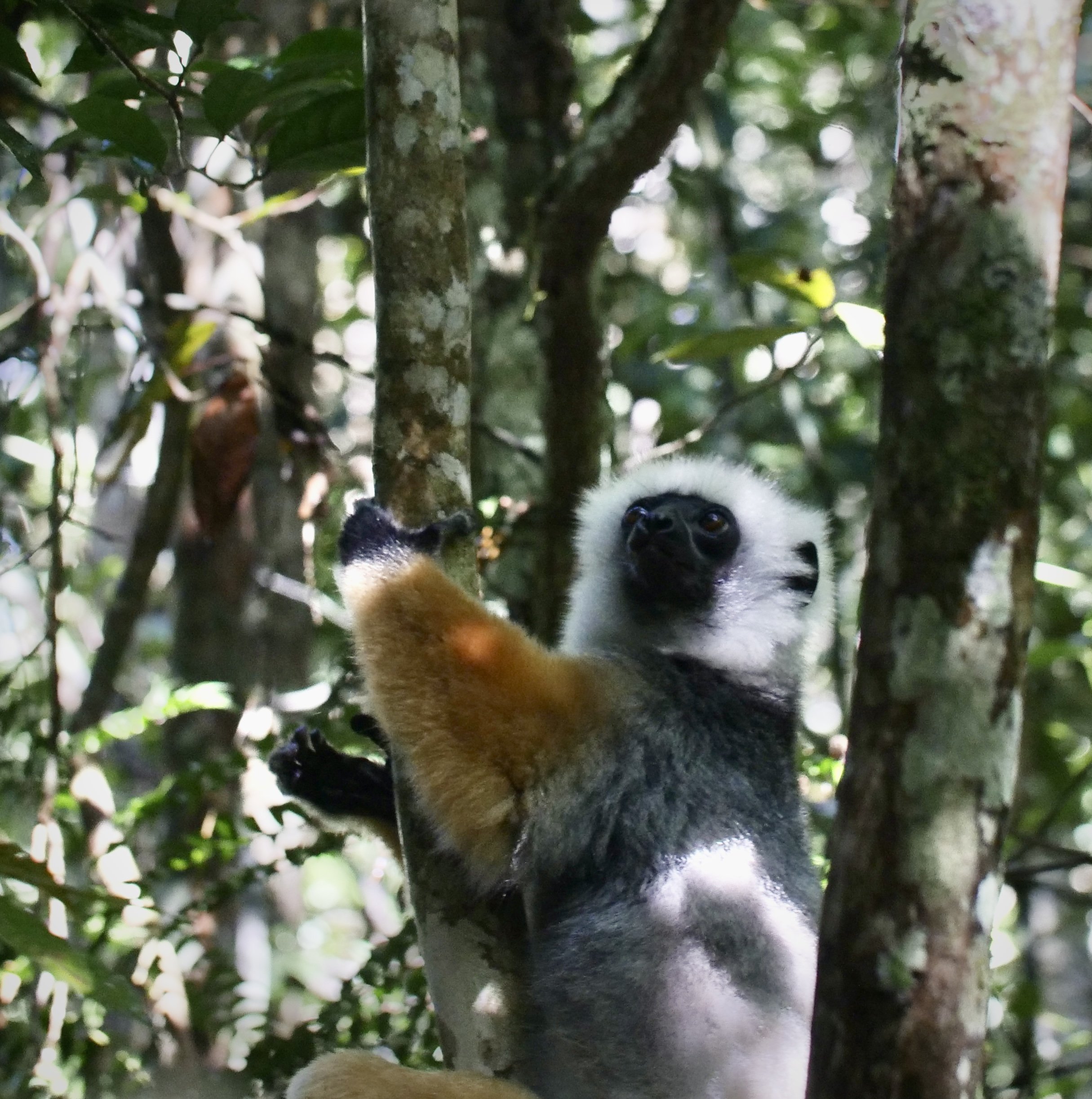
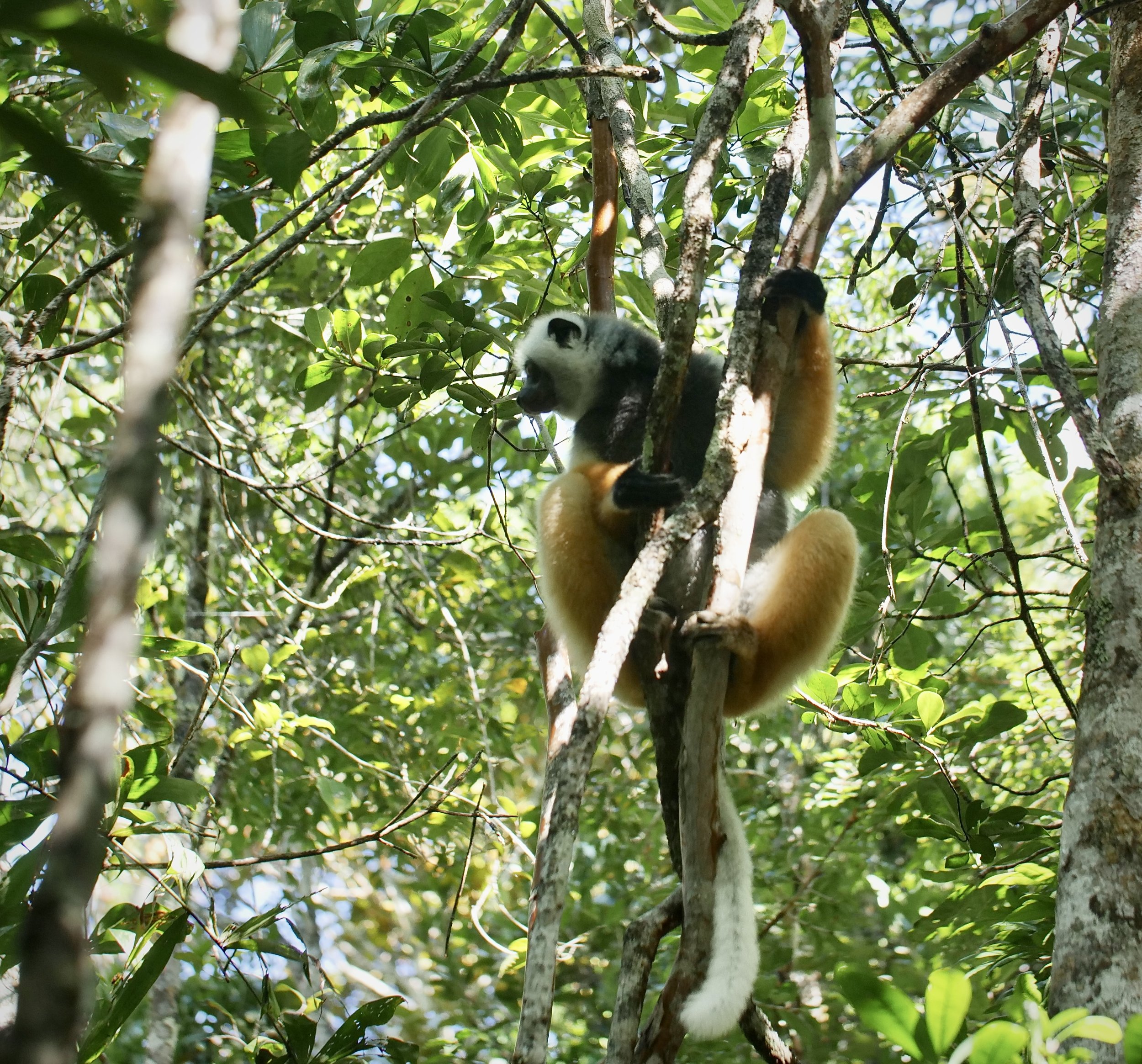
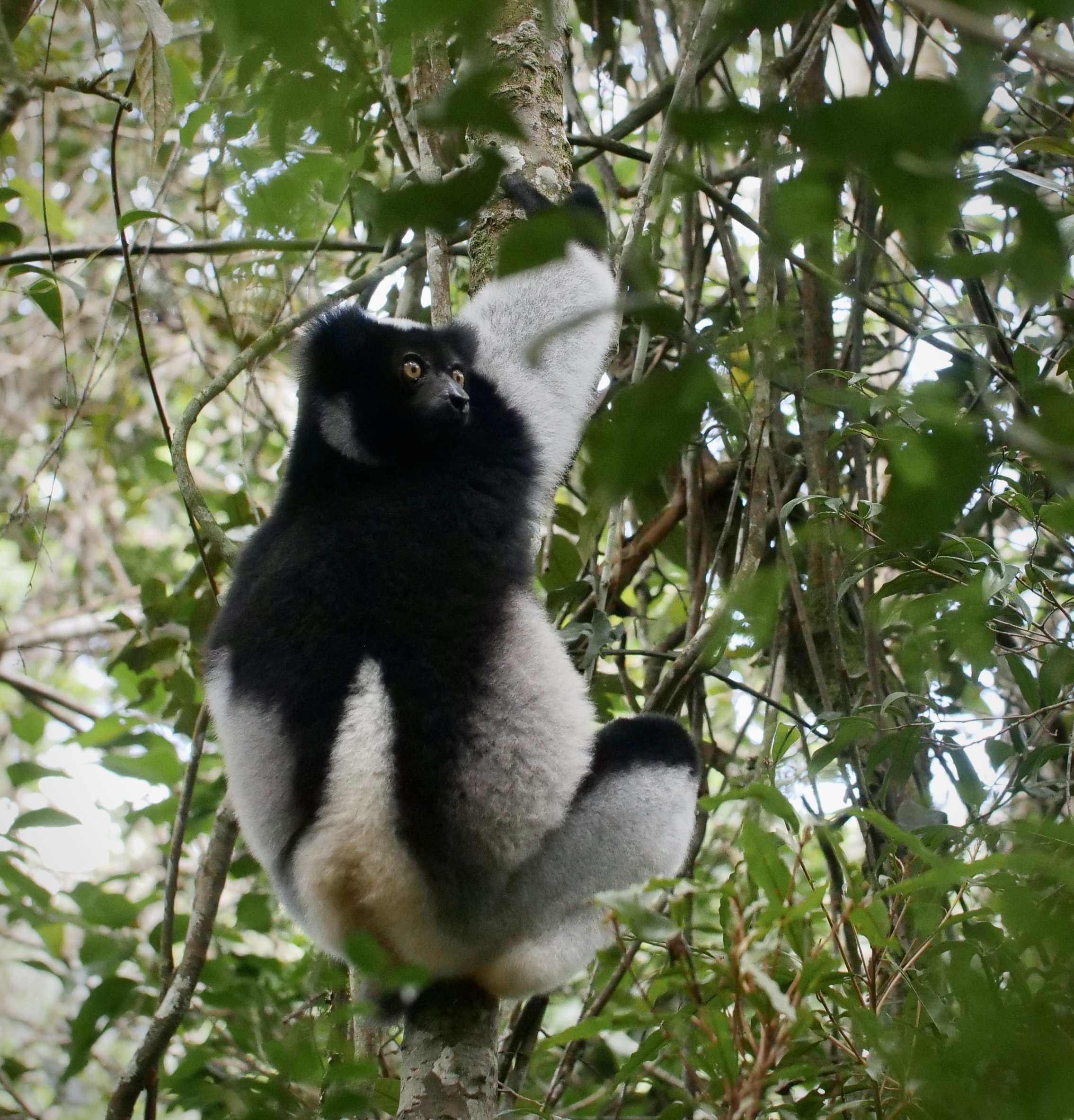






The diademed sifaka, Propithecus diadema, an endangered species of sifaka, one of the lemurs endemic to rainforests in eastern Madagascar, followed by the Indri, Indri indri, one of the largest living lemurs.
Sounds on! Listen to the territorial call of the Indri
We spent four full days exploring the rainforest in and around Andasibe-Mantadia National Park. We hiked along the Tsakoka and Rianasoa trails and the protected area of Vohimana, an off-the-beaten-track reserve where we photographed species usually not found at Andasibe.
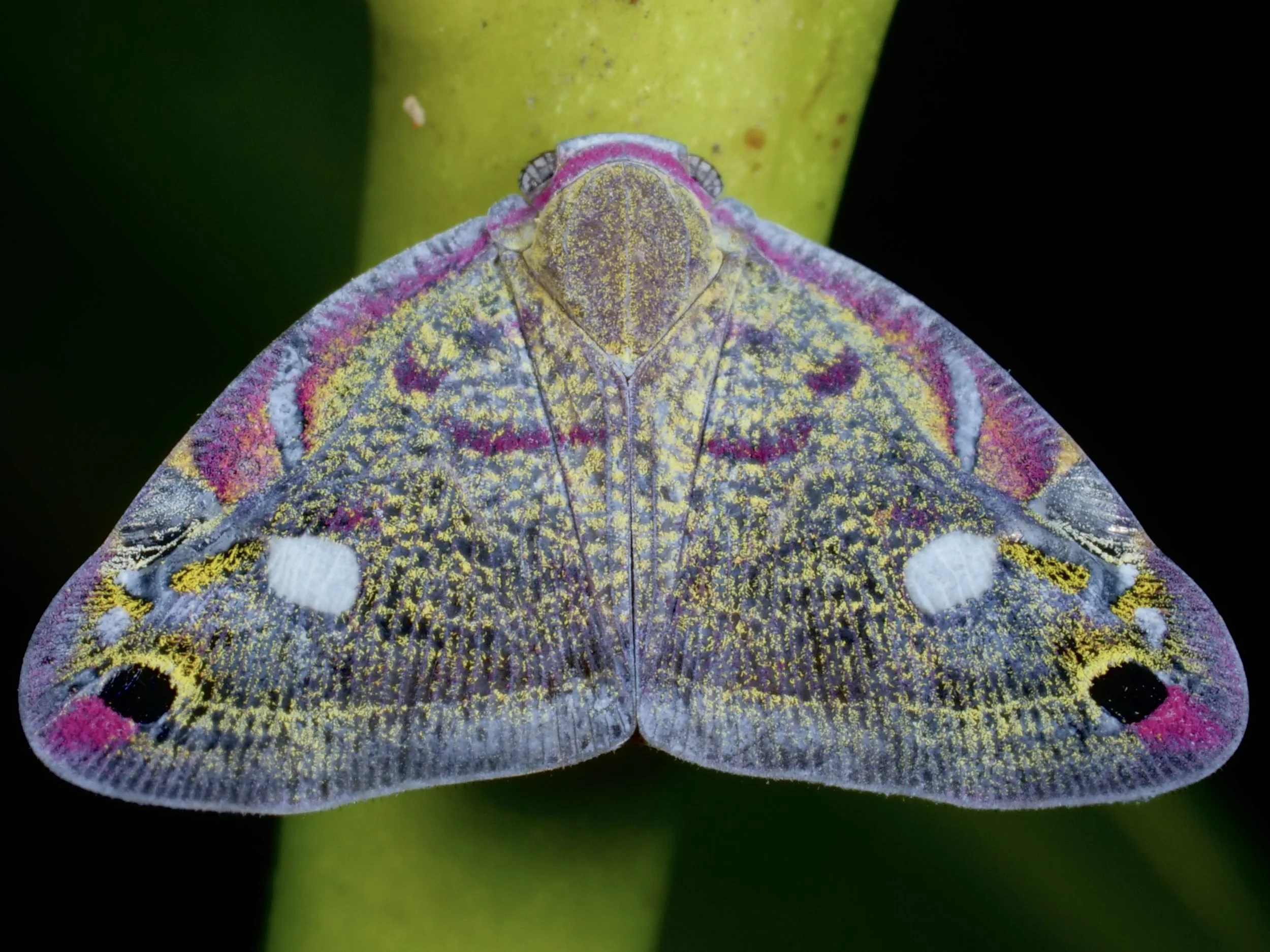
I mistook this planthopper for a colorful moth at first - those wings are 🔥
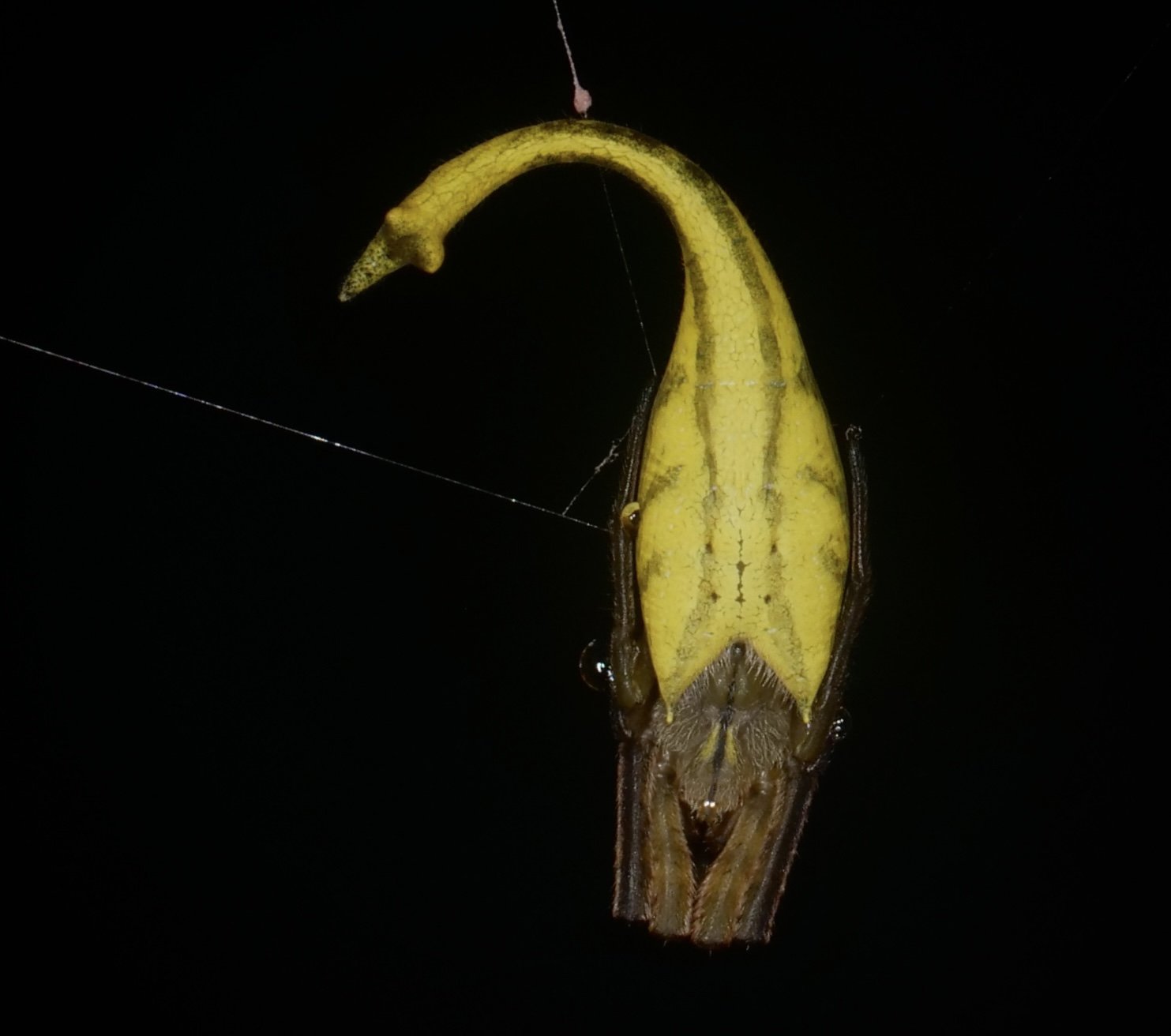
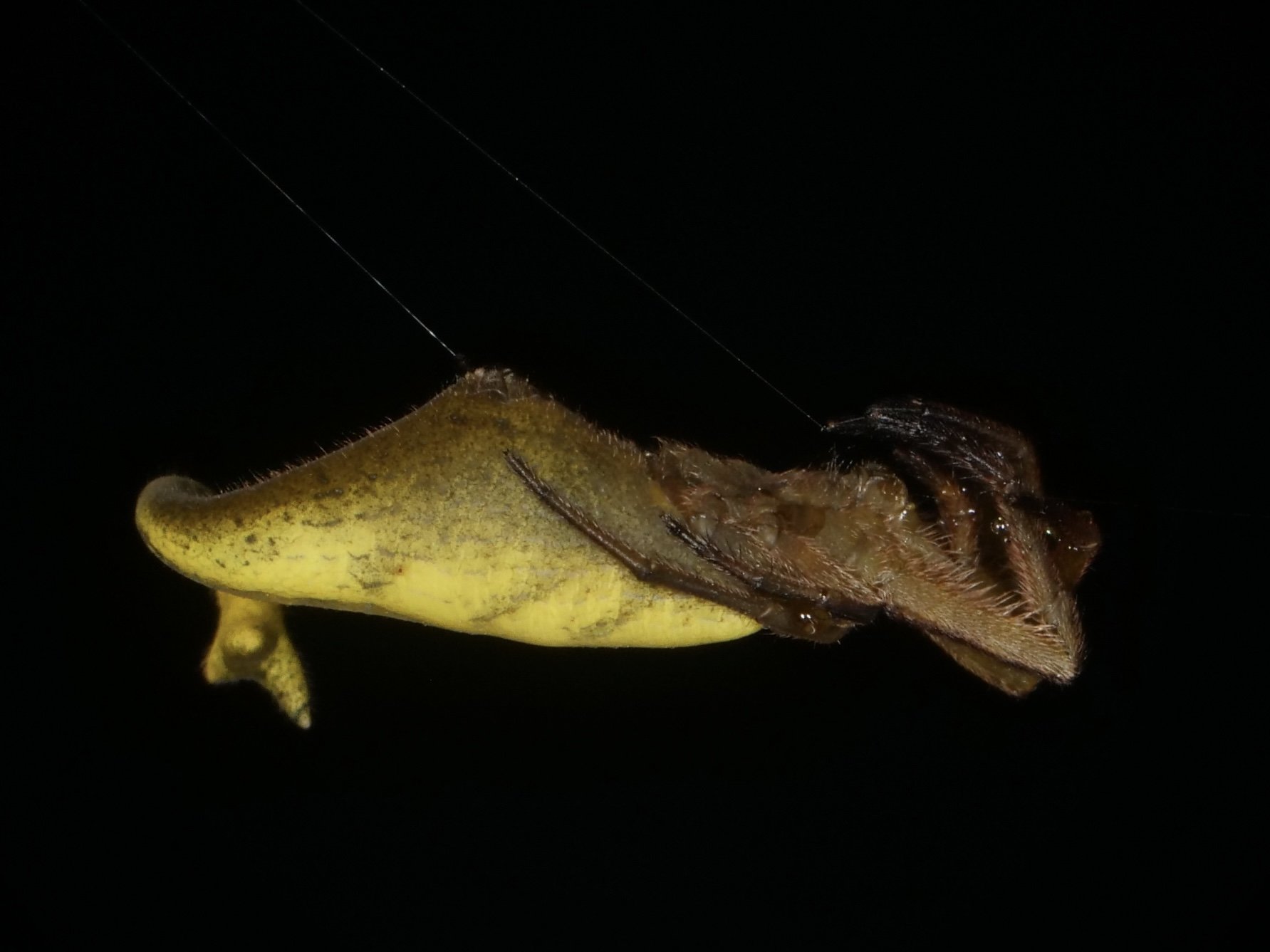


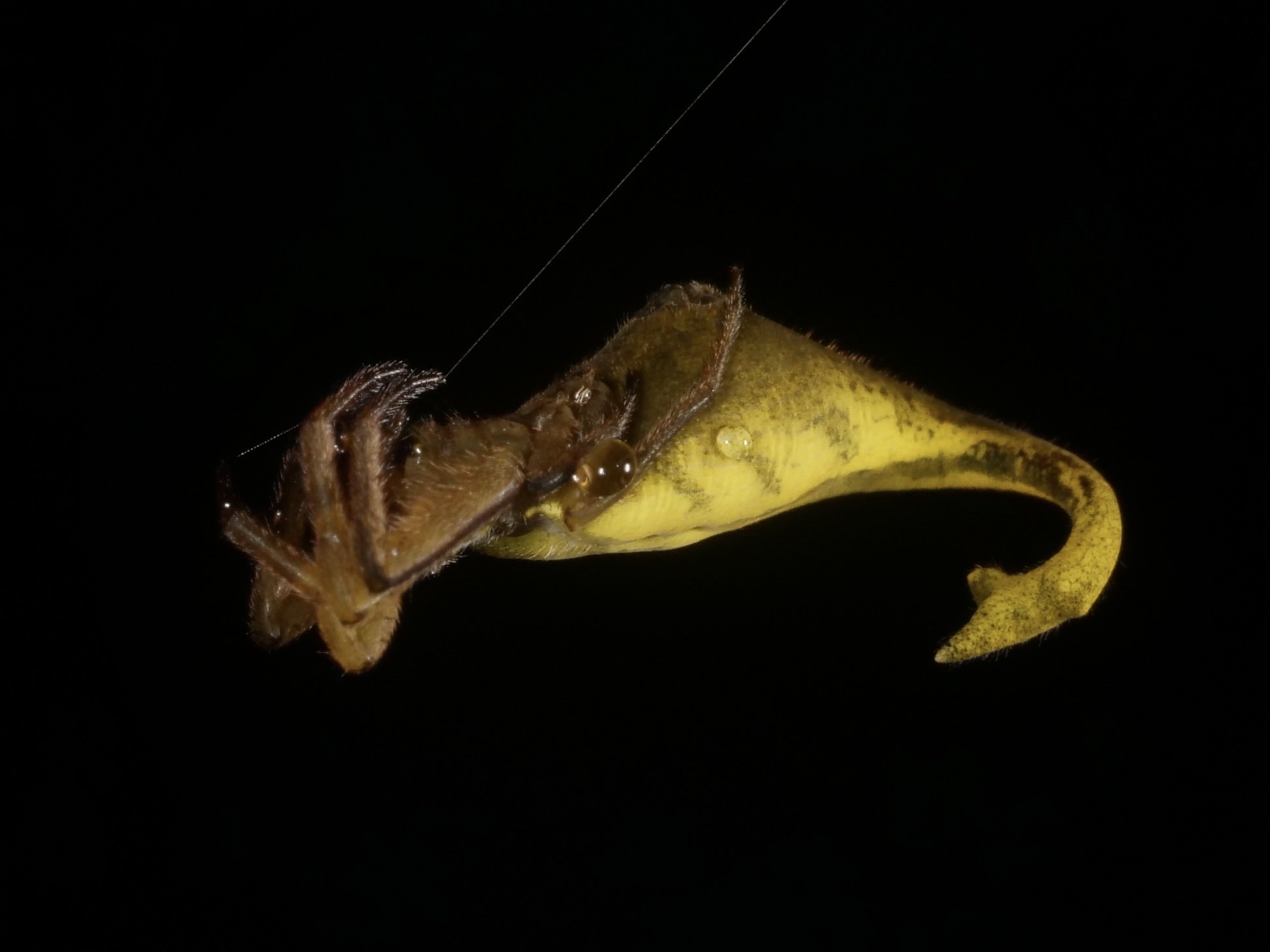
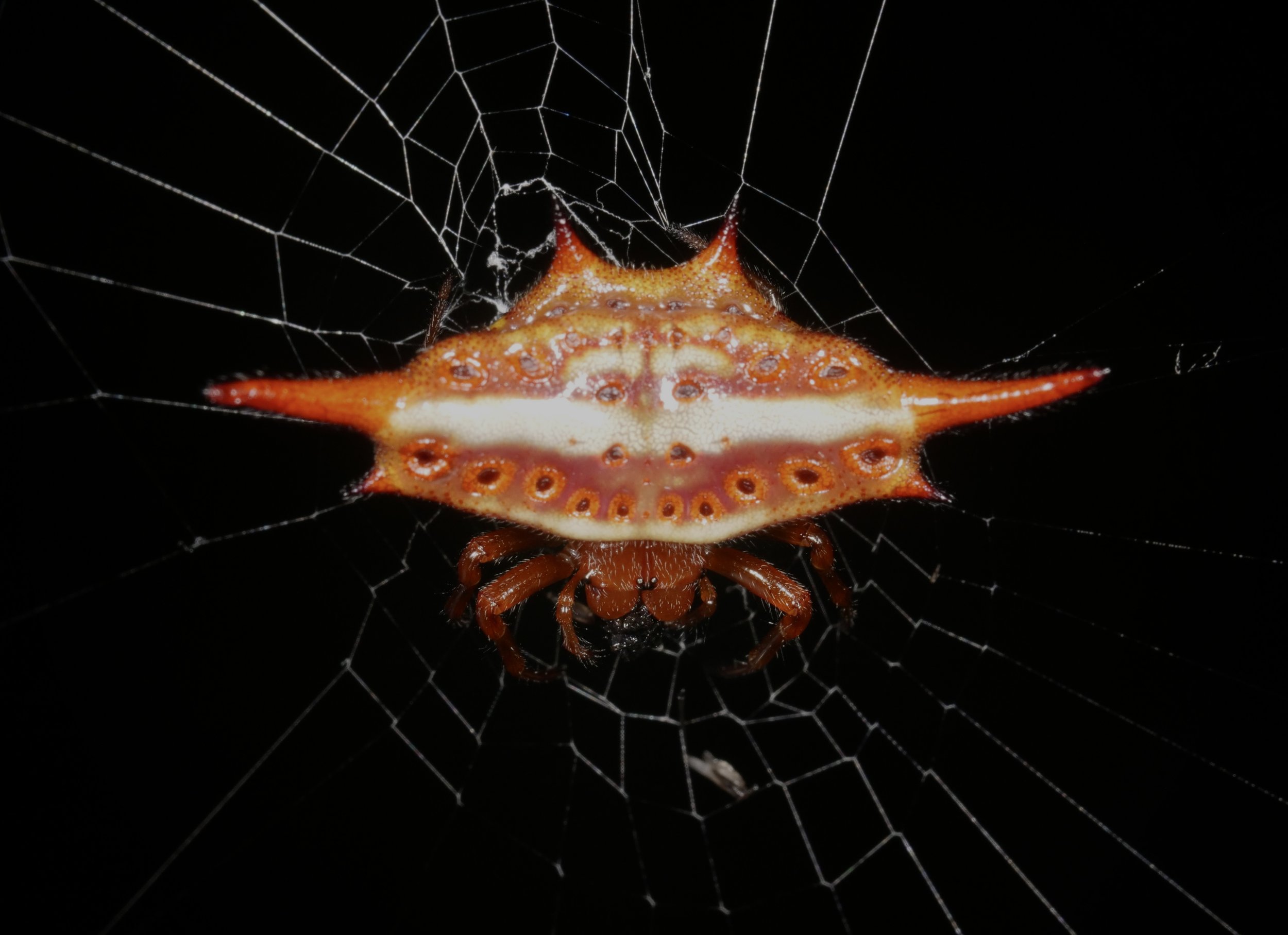
One of my favorite arachnid finds in Madagascar, the scorpion-tailed spider (Arachnura genus). I would have guessed the tail-like structure could have evolved as a modified spinneret organ but looks to be a completely separate appendage. Video at the end shows a little tail-curl. Presumably used to deter would-be parasitoids?
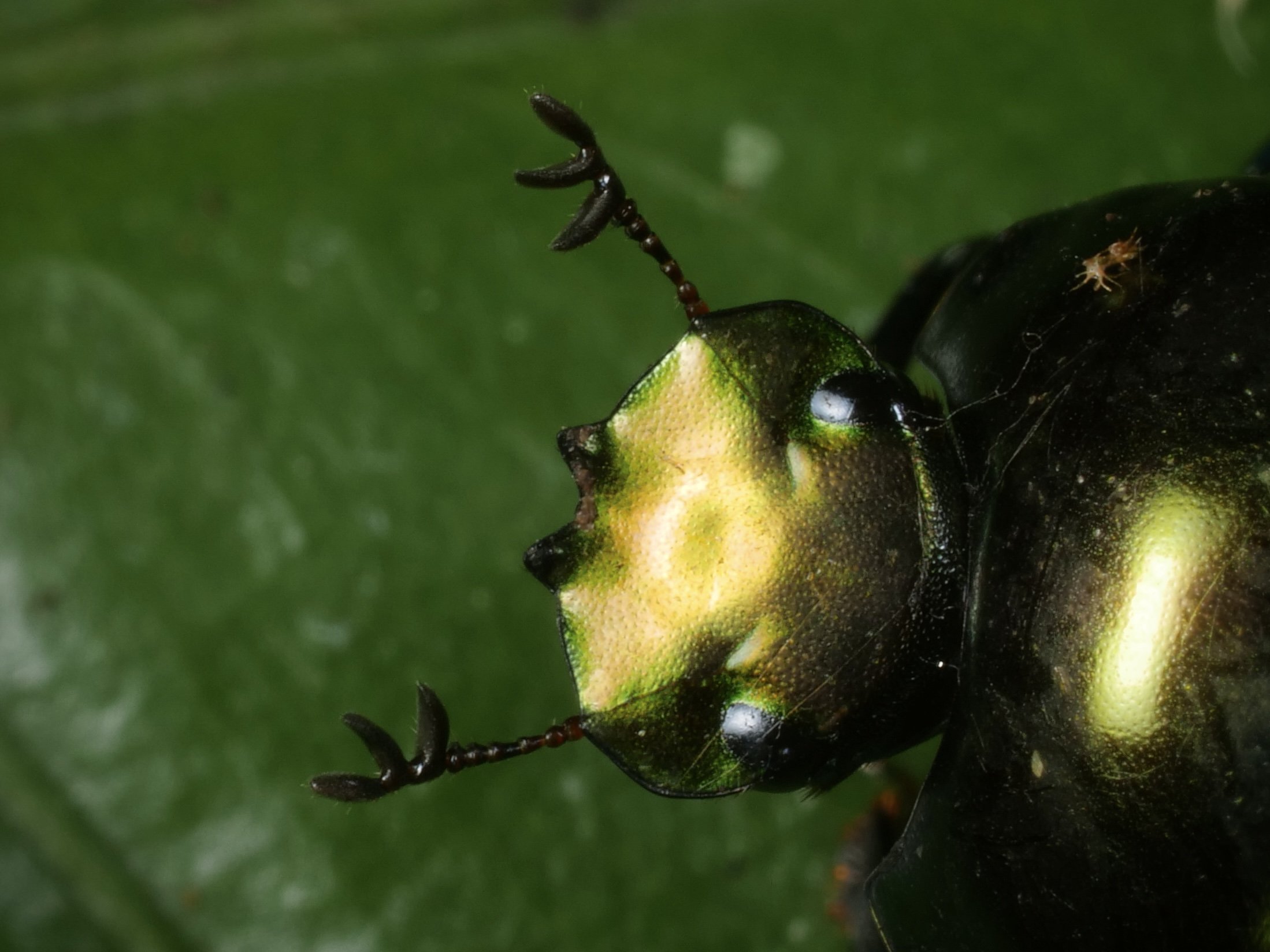
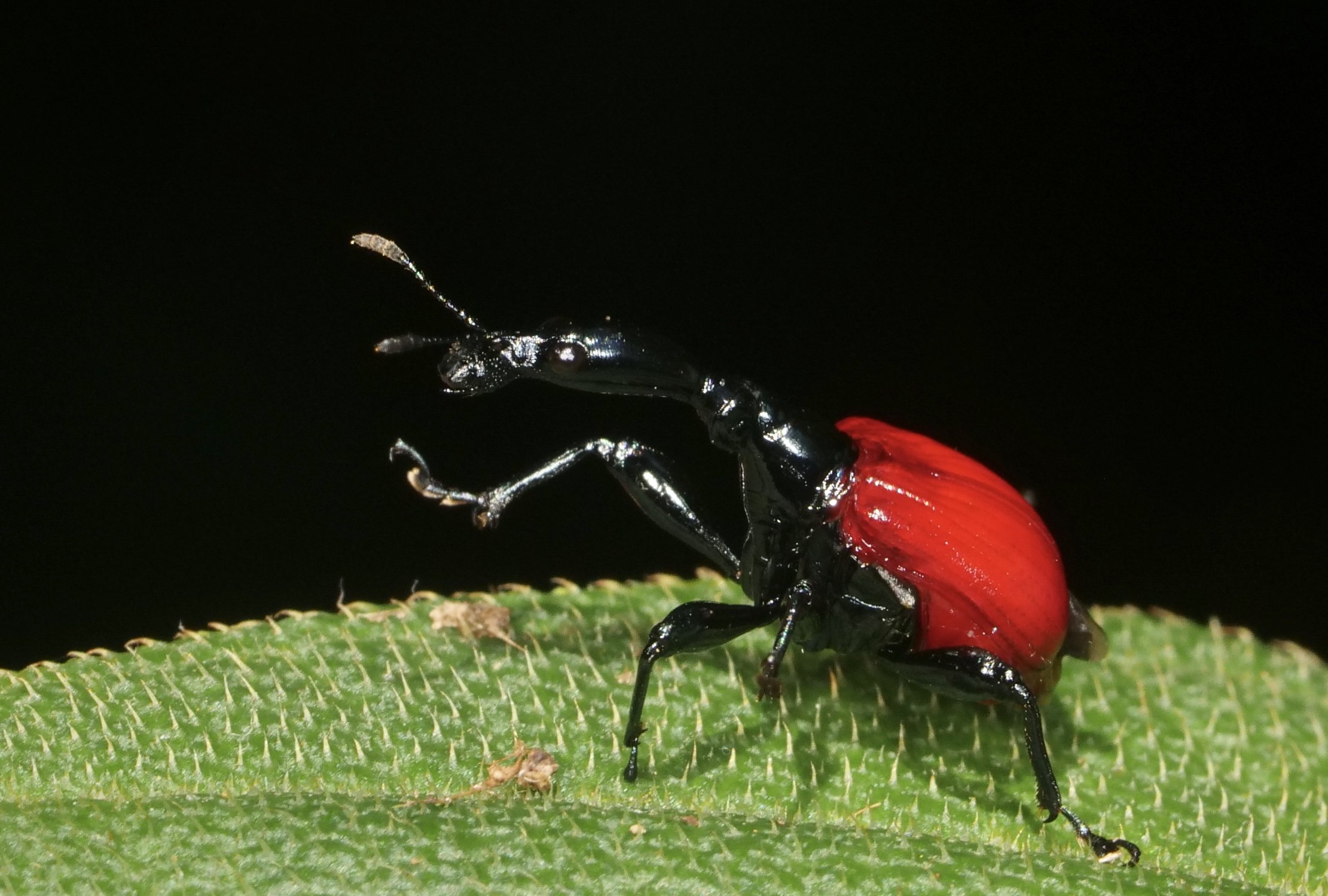
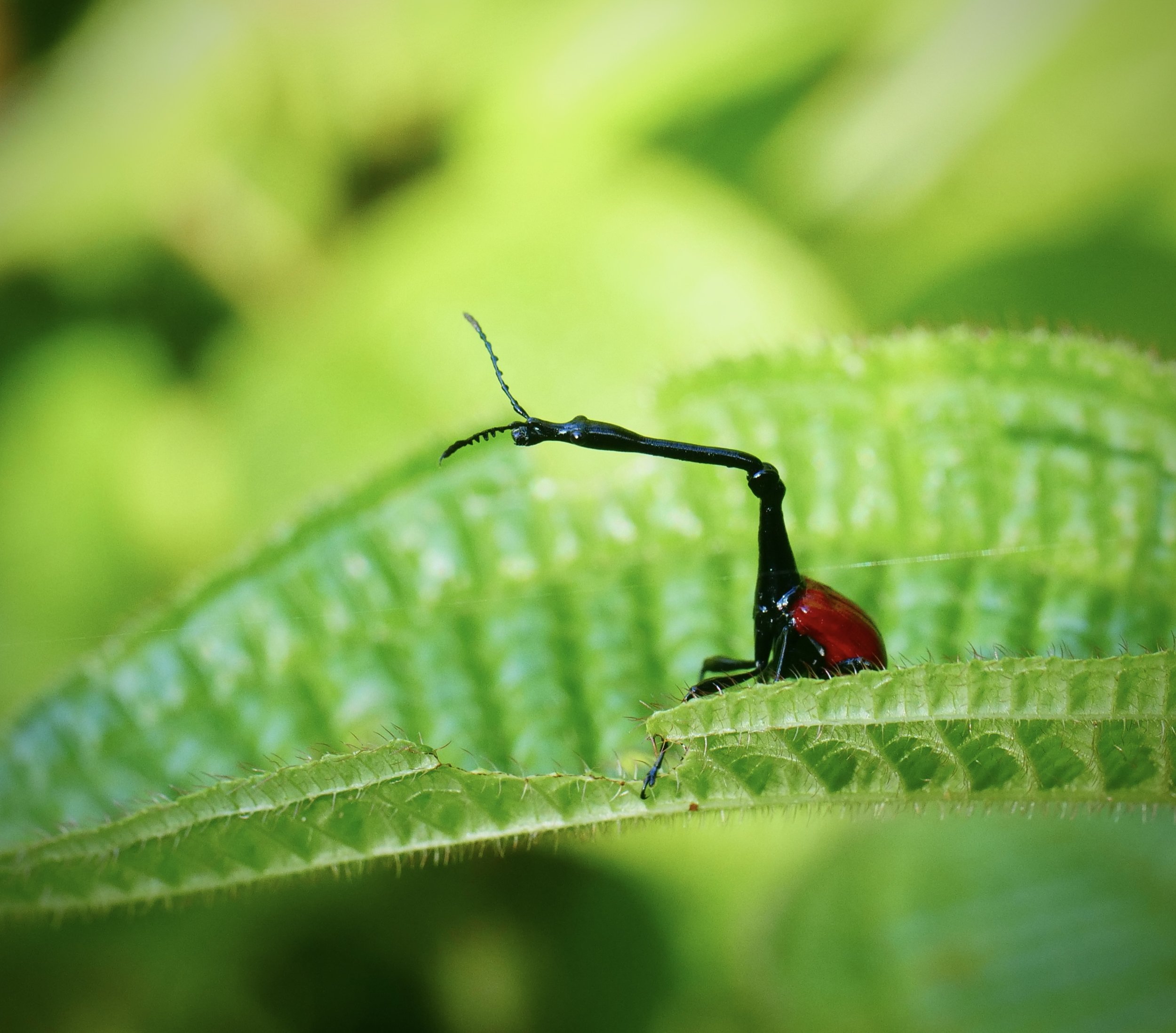
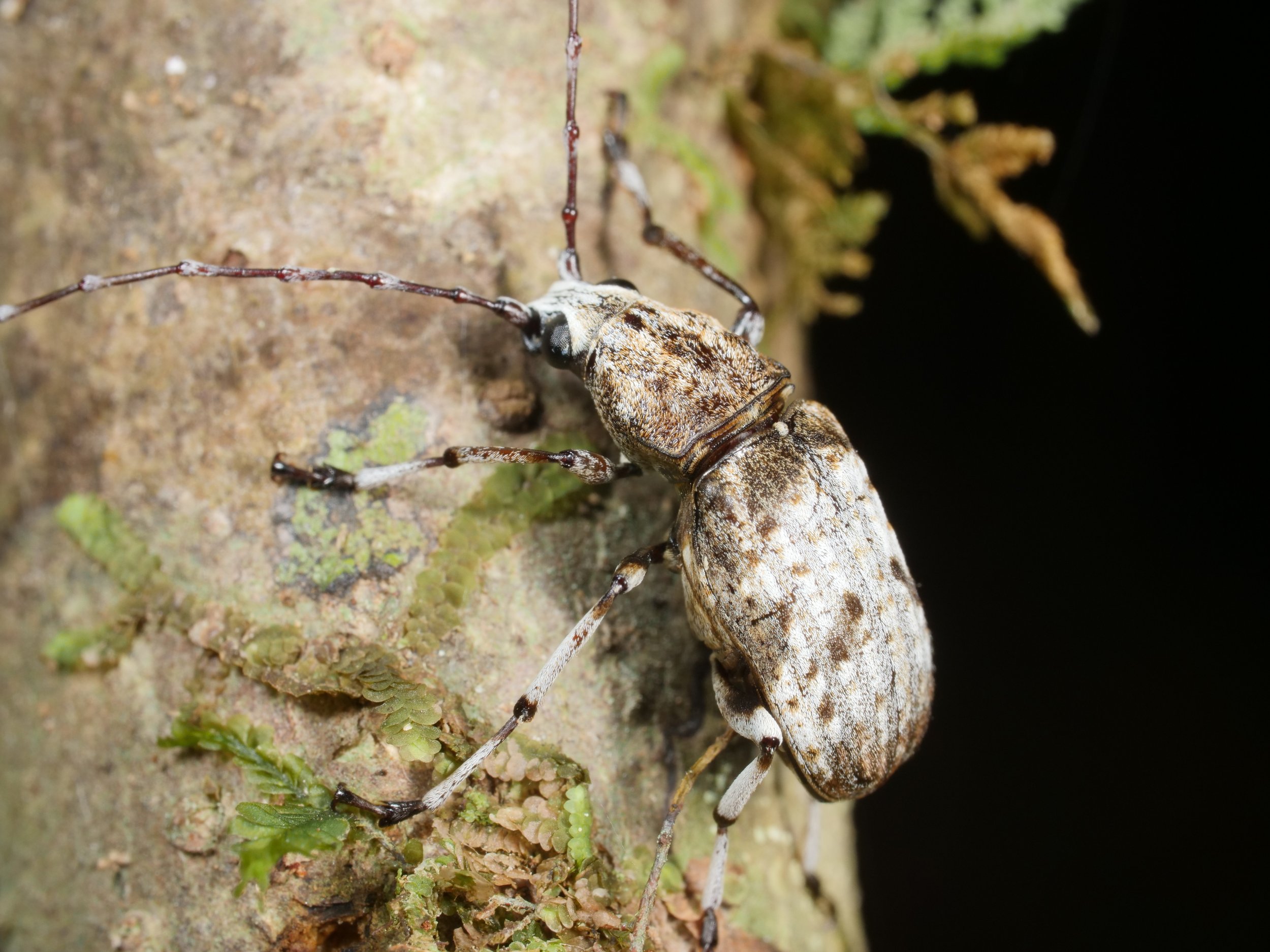
The giraffe weevil Trachelophorus giraffa is a weevil found across parts of Southern Africa, India, and southeast Asia. As the name implies, this species has a peculiar extended neck, much like that of a giraffe. It is much more pronounced in the male (right) compared to the female (left).
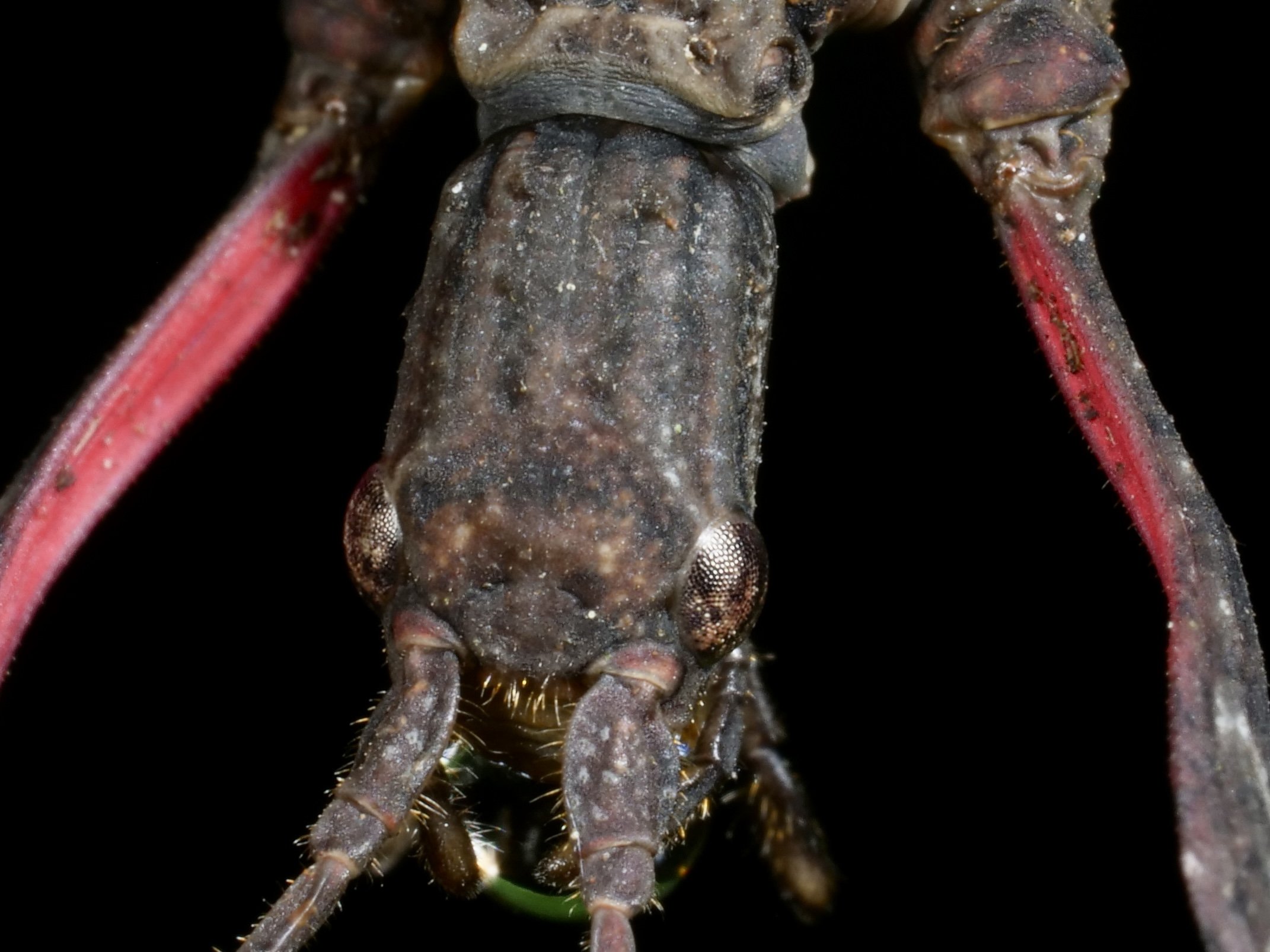
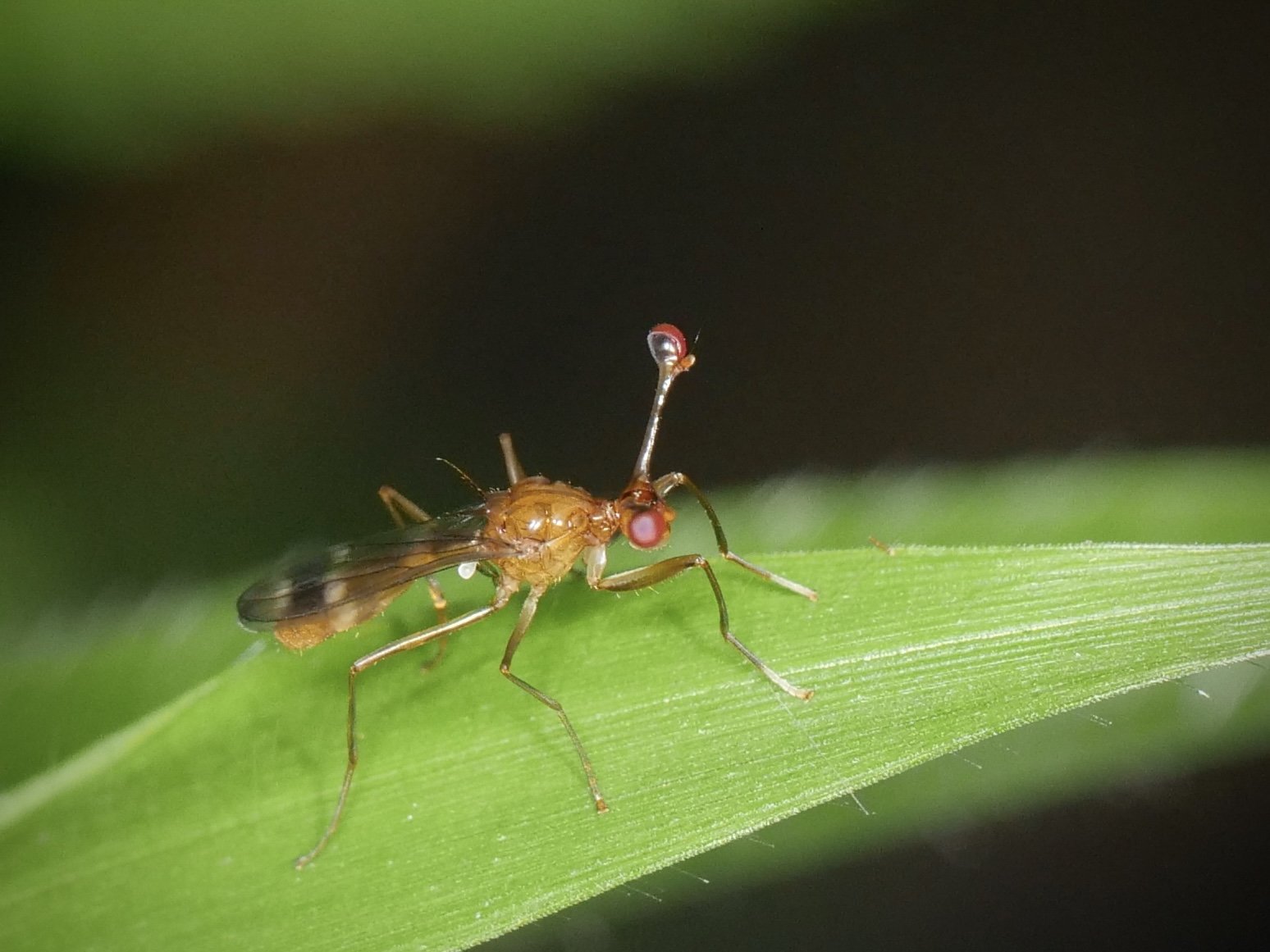
The stalk-eyed fly showcases another bizarre case of sexual dimorphism. Belonging to the family Diopsidae, members possess "eyestalks", long projections from sides of the head with eyes at the end.
We also hiked the poorly-explored Iaroka forest and visited Mitsinjo, a community-run reserve known for its incredible diversity of frog species. During the nights, we explored the trails at Mitsinjo and Analamazaotra reserves.
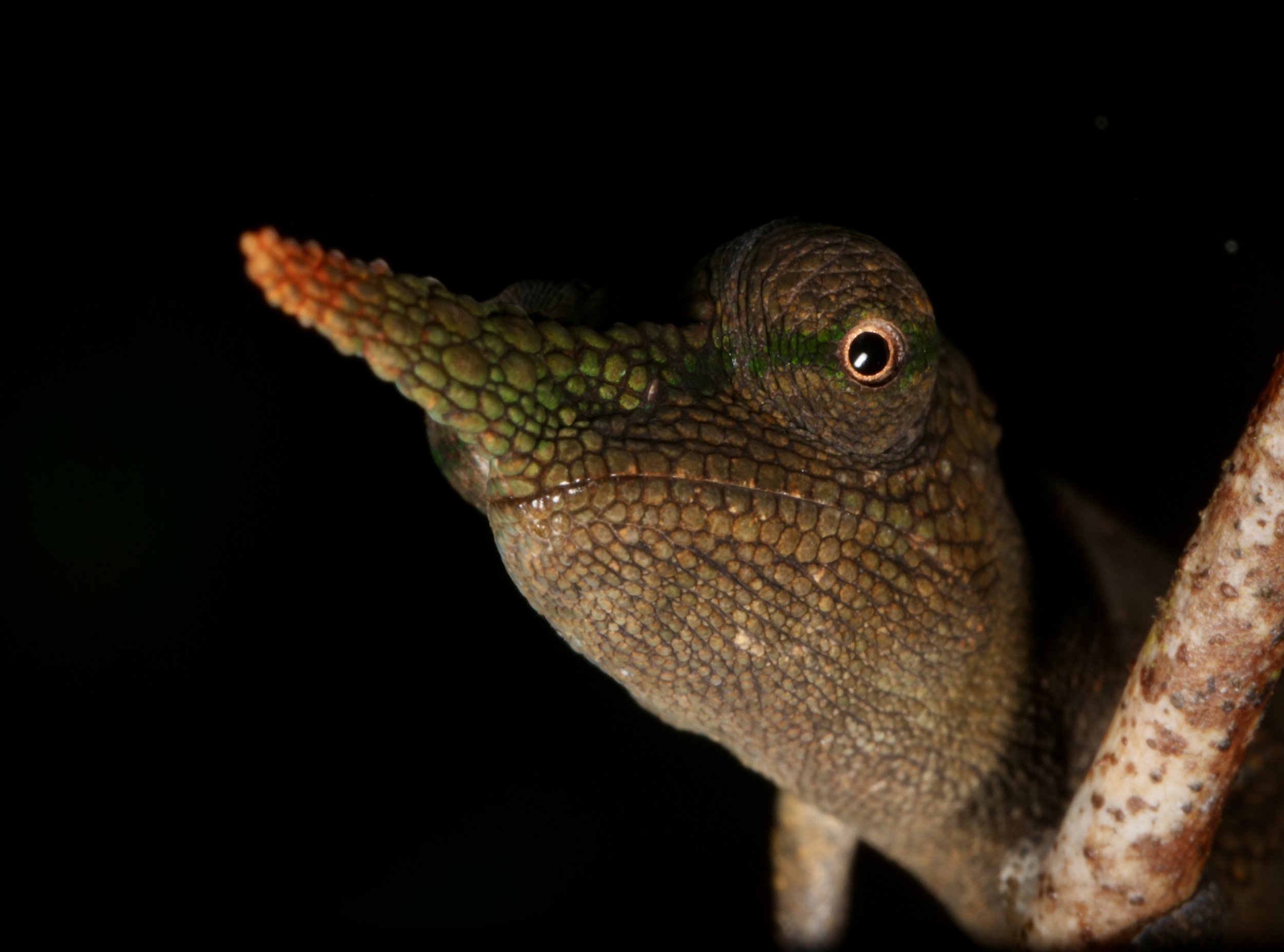
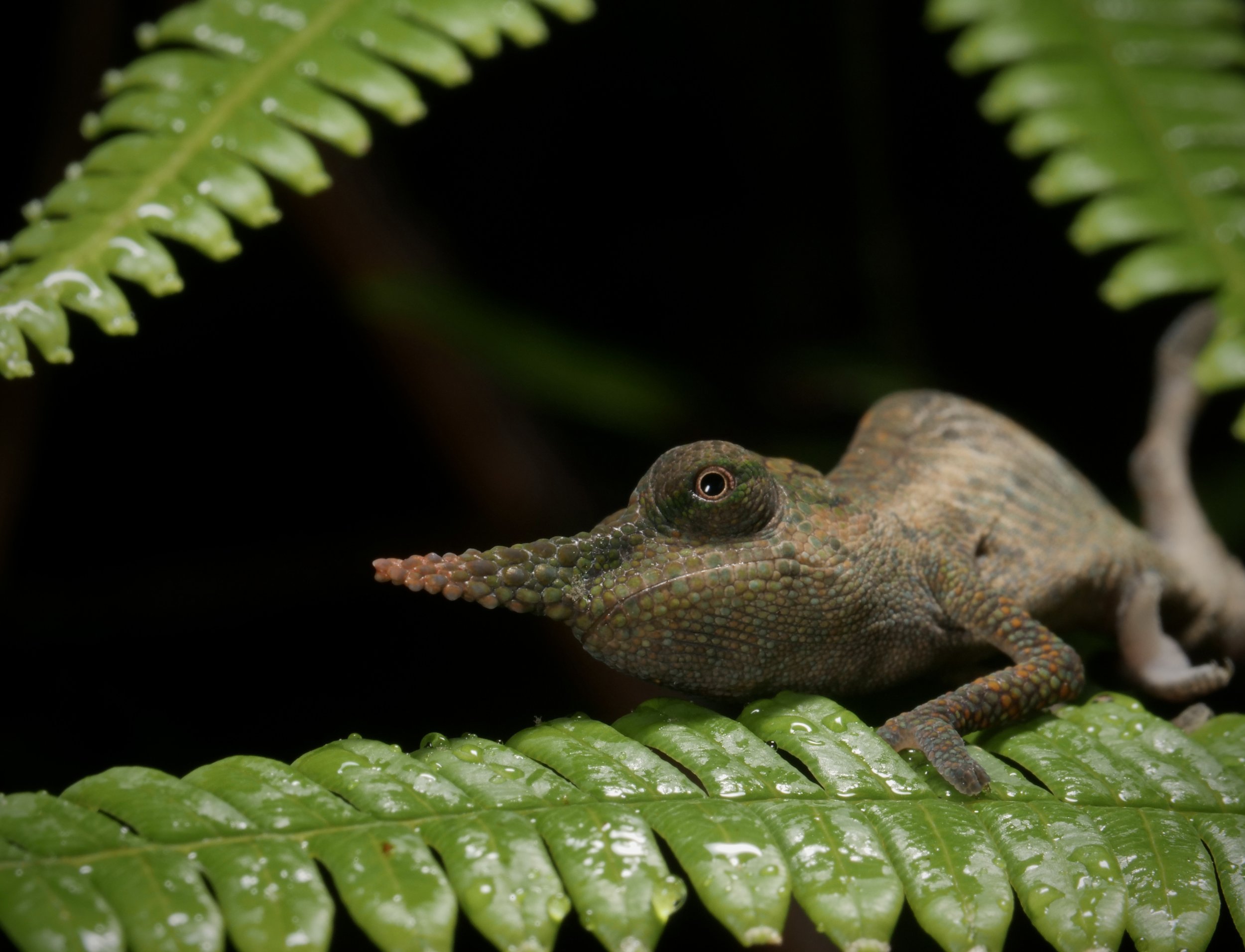
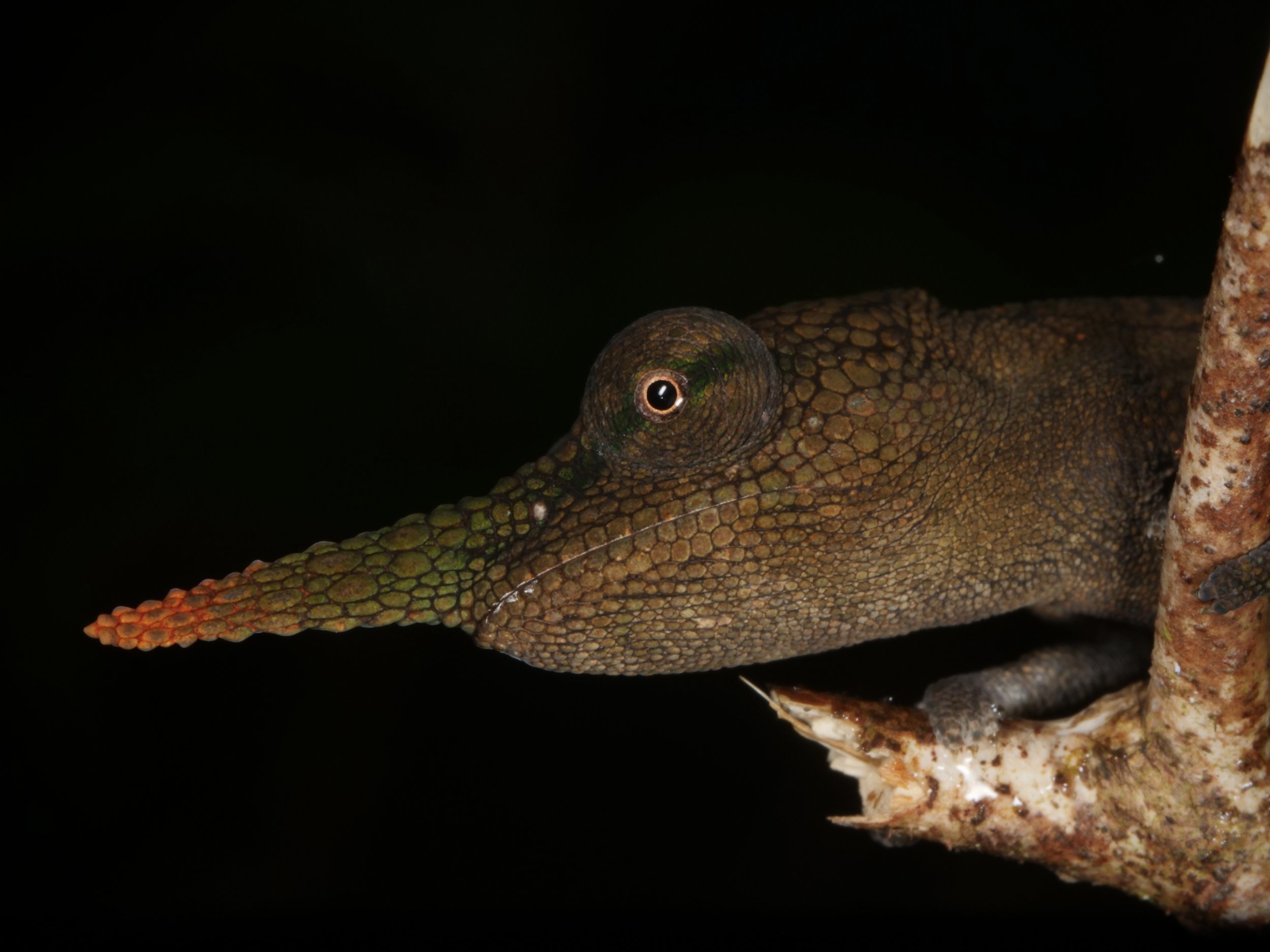
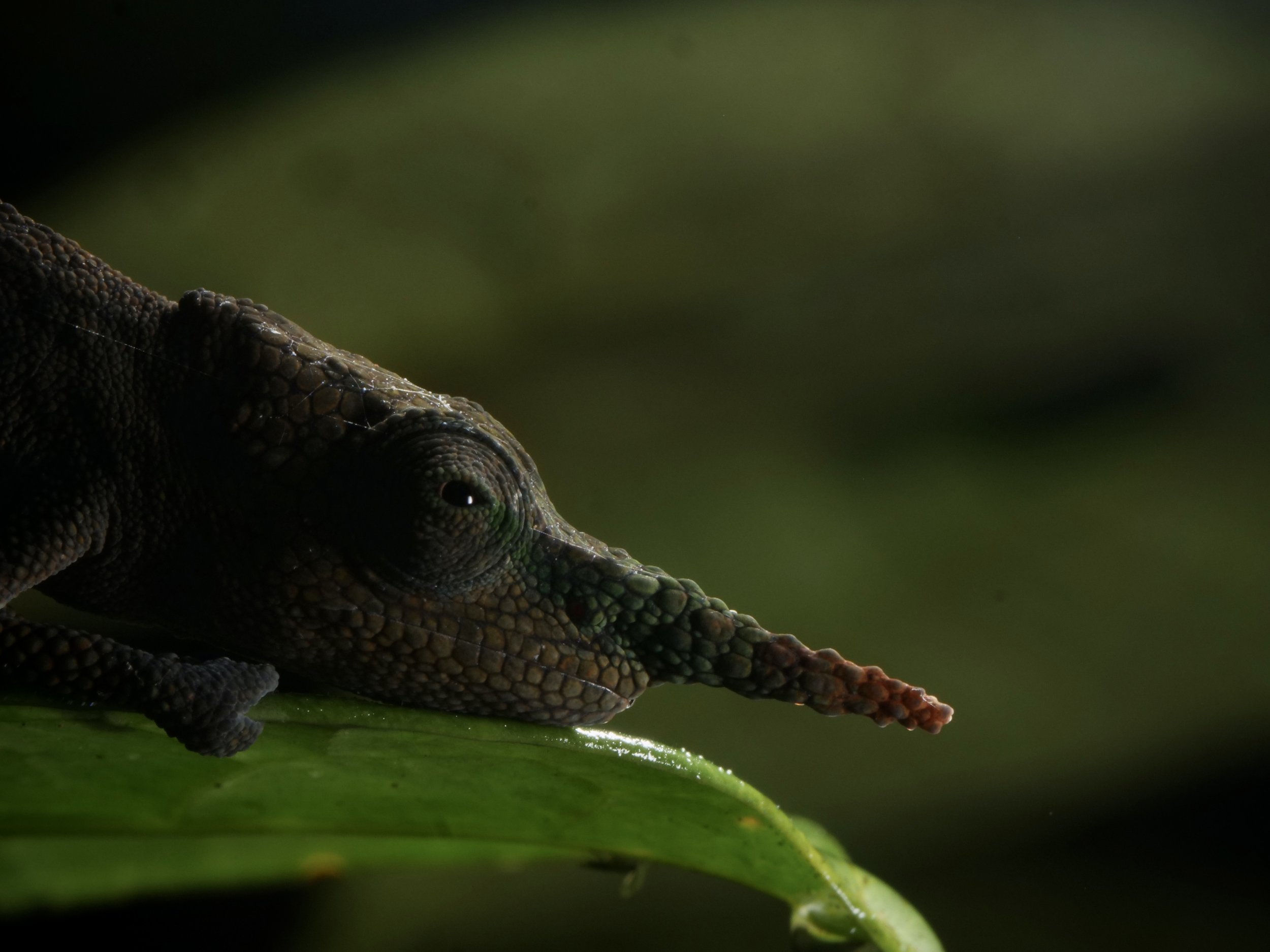




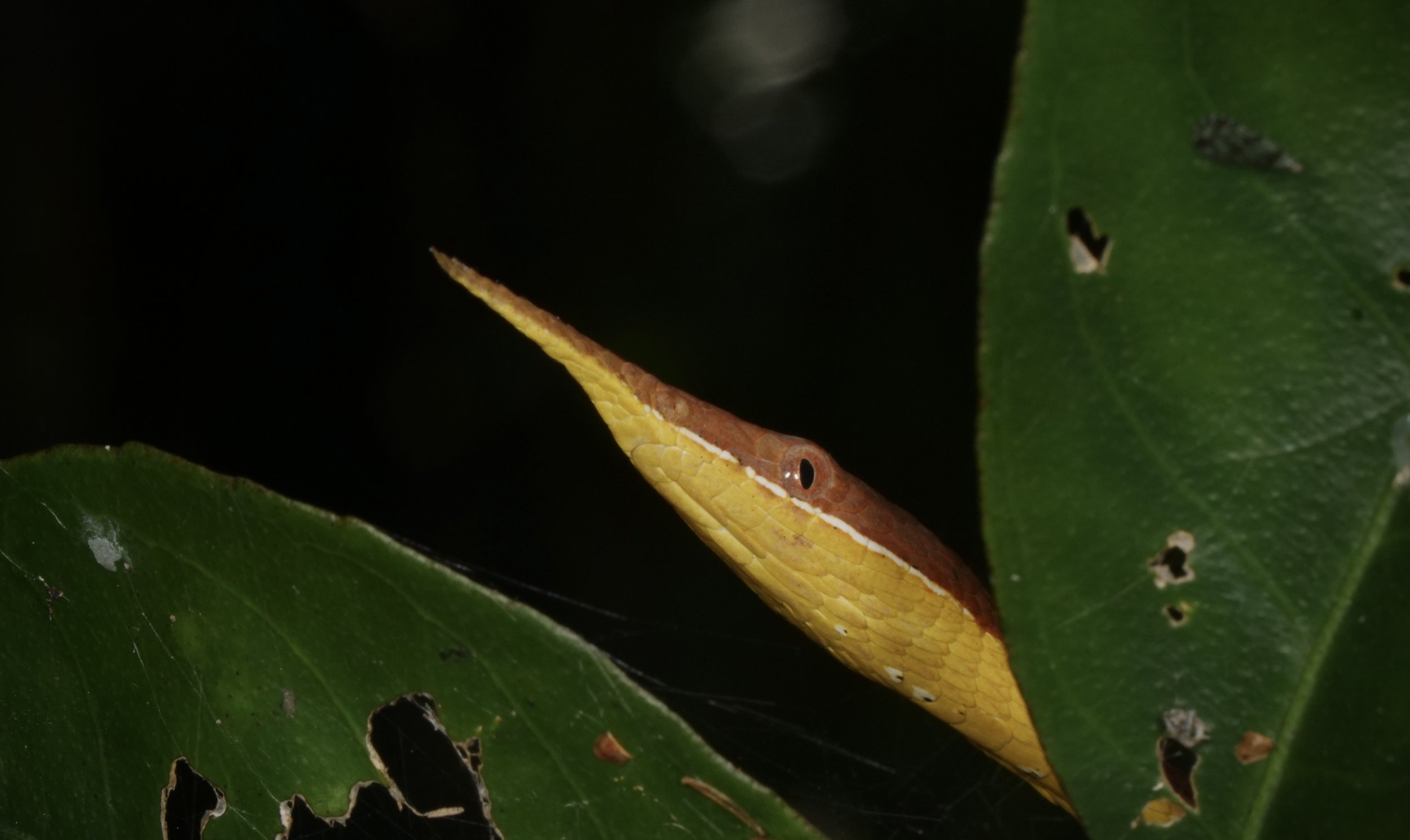
Vohimana forest in Madagascar, home to the bizarre and beautiful ‘Lance-nose’ chameleon Calumma gallus
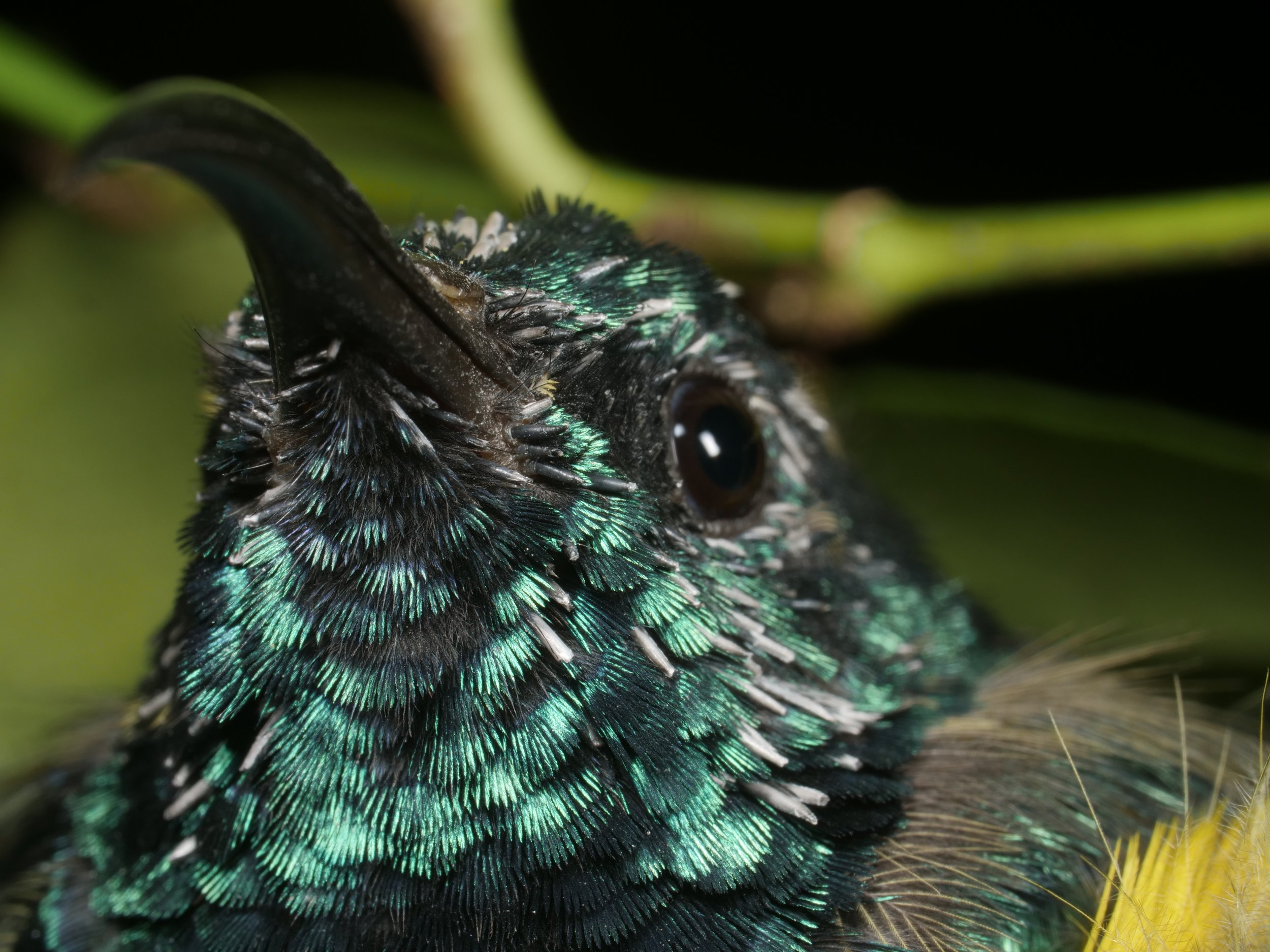
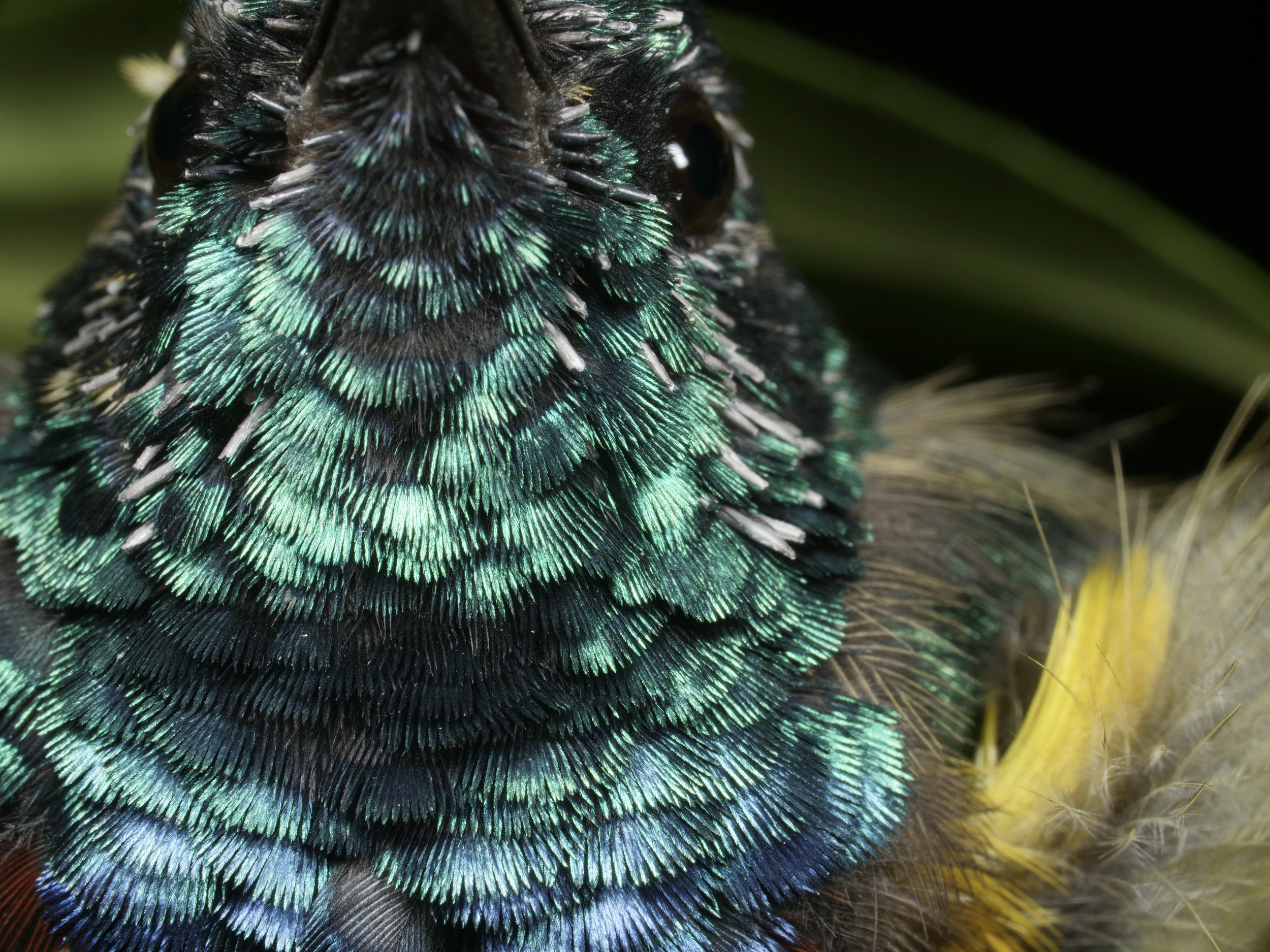
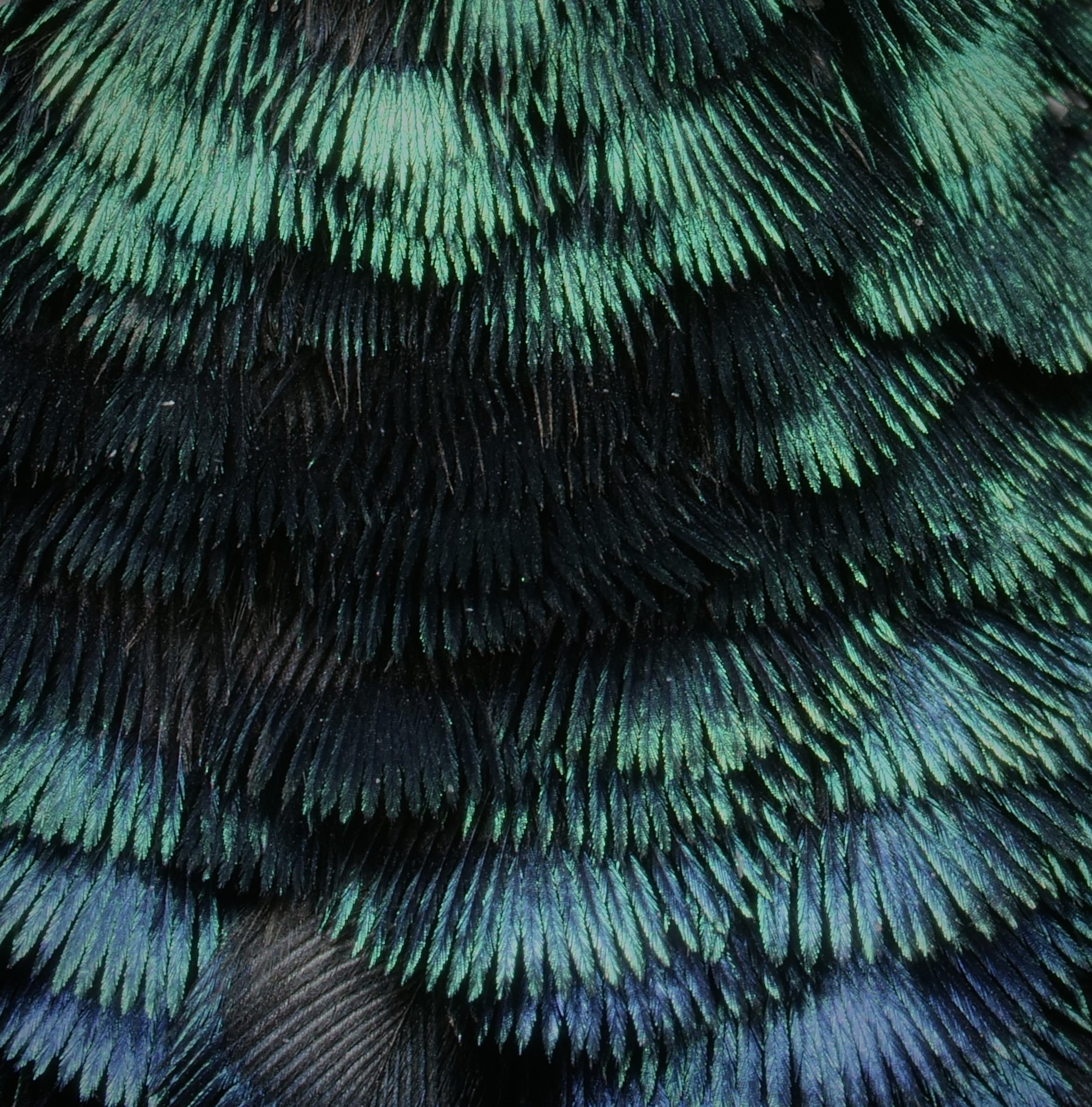



Beautiful iridescent feathers of a sunbird
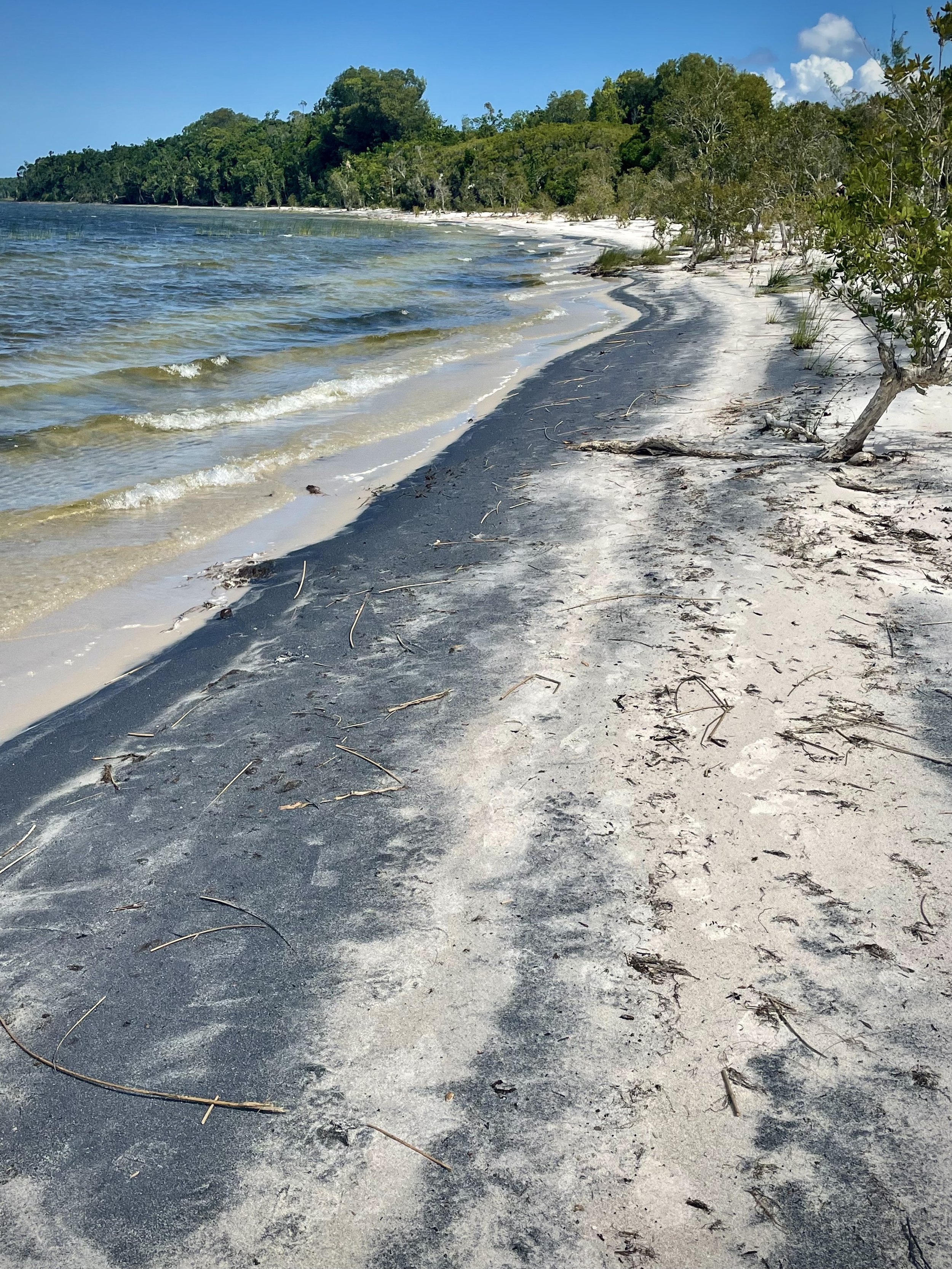
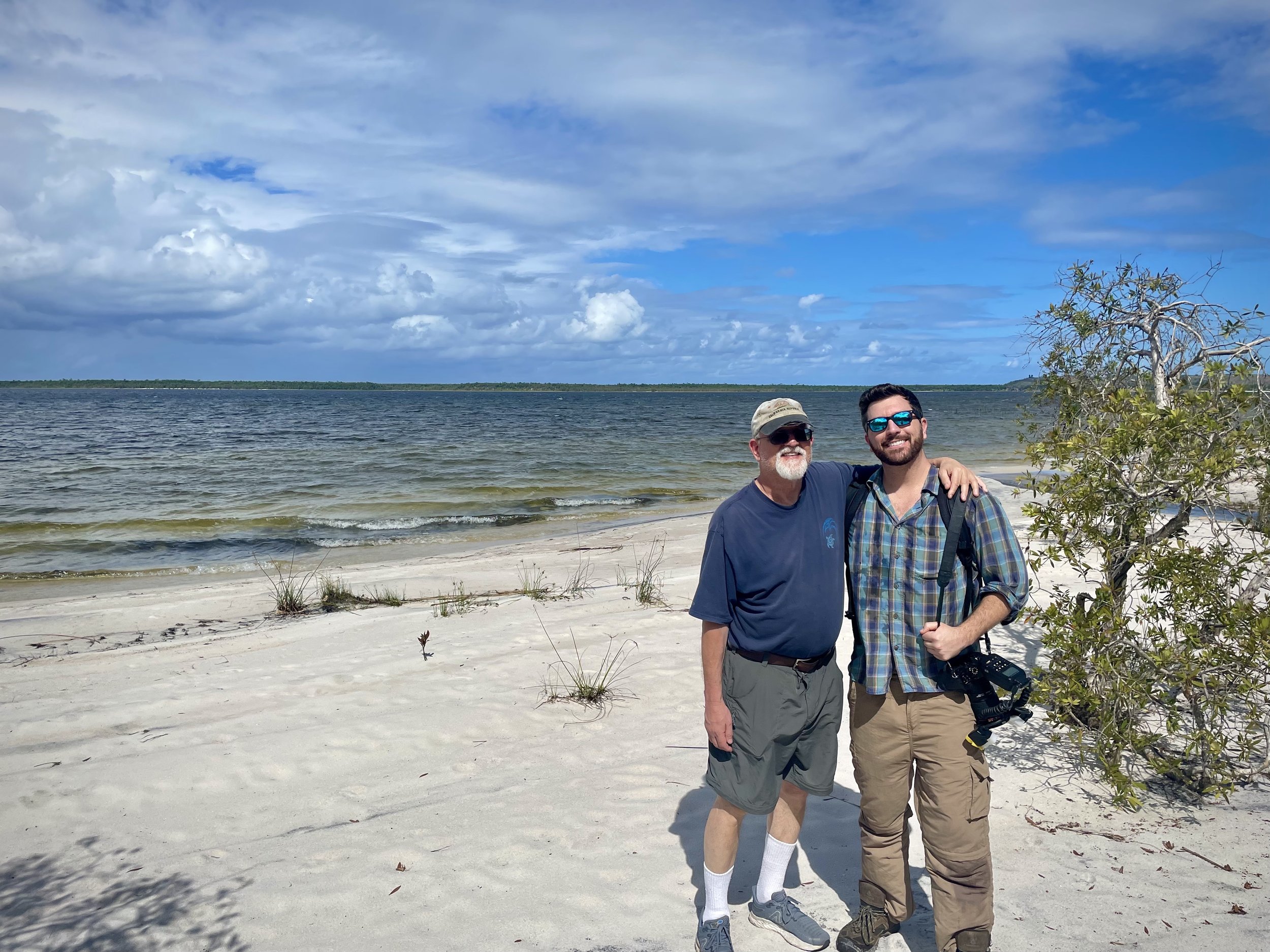
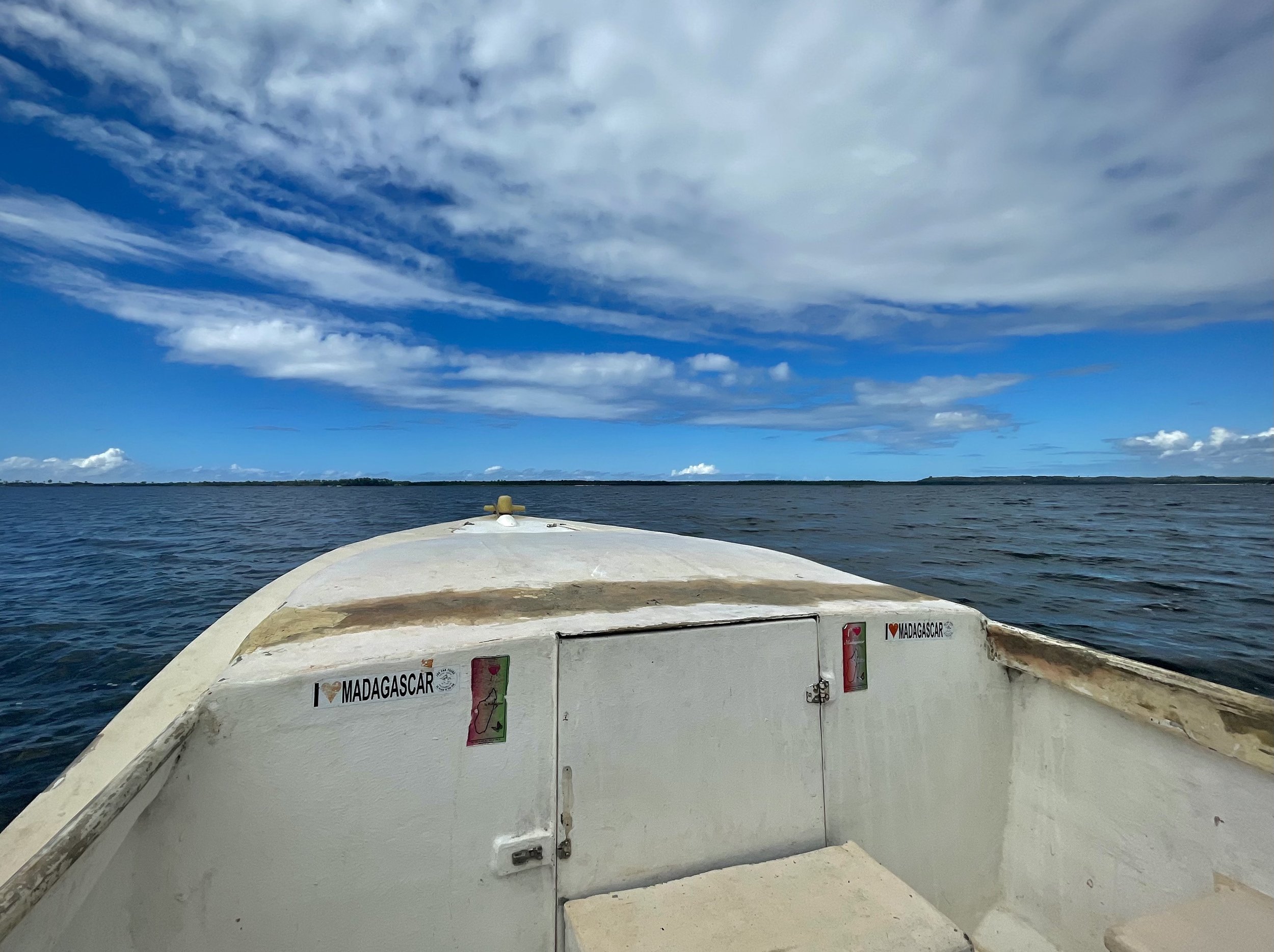
Finally, we set out on a long drive from Andasibe to a two- hour motorized canoe ride which took us to Ankanin’ny Nofy, a beautiful coastline site in a relaxing shoreline atmosphere and stayed at hotel Palmarium.
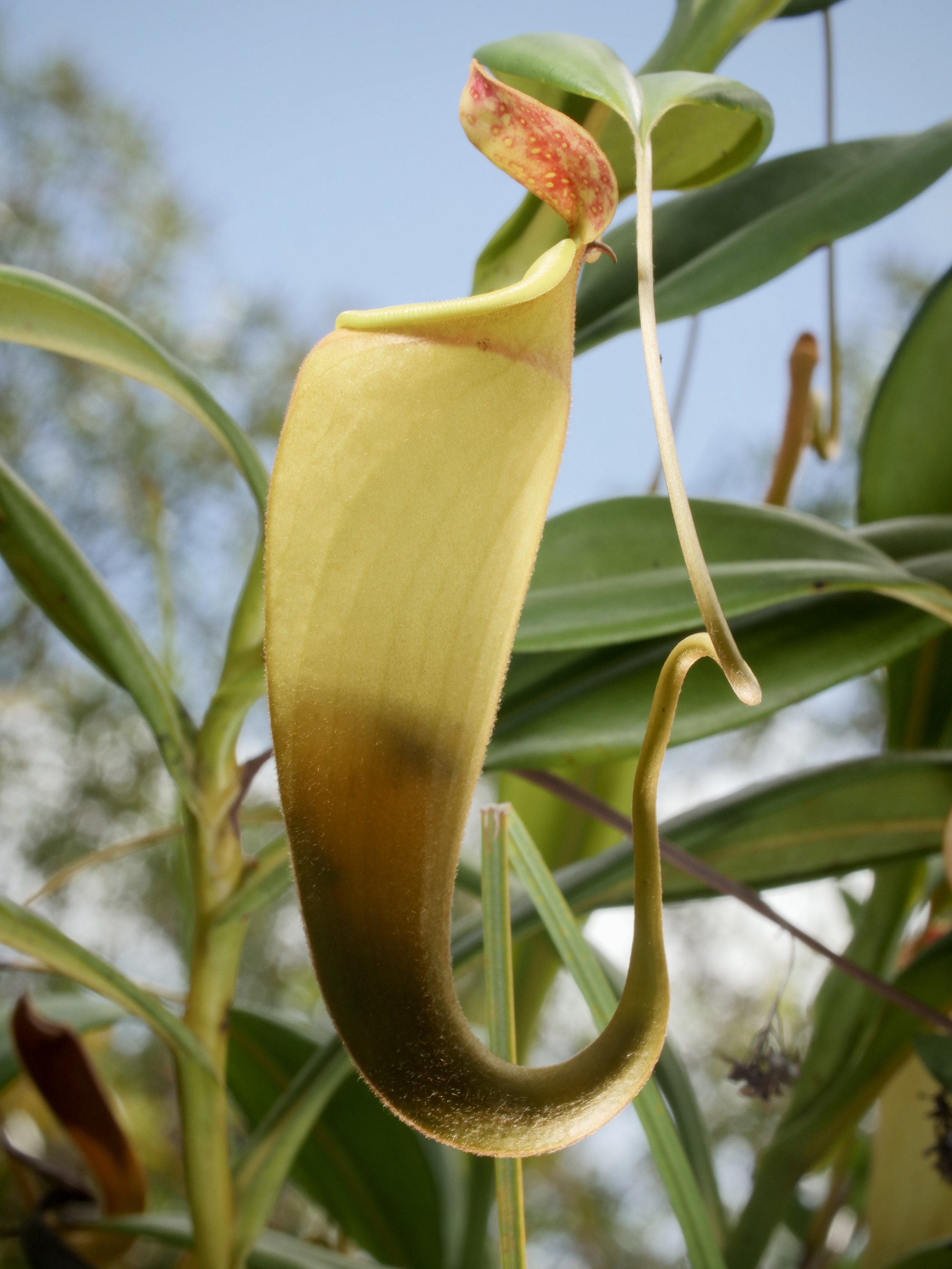
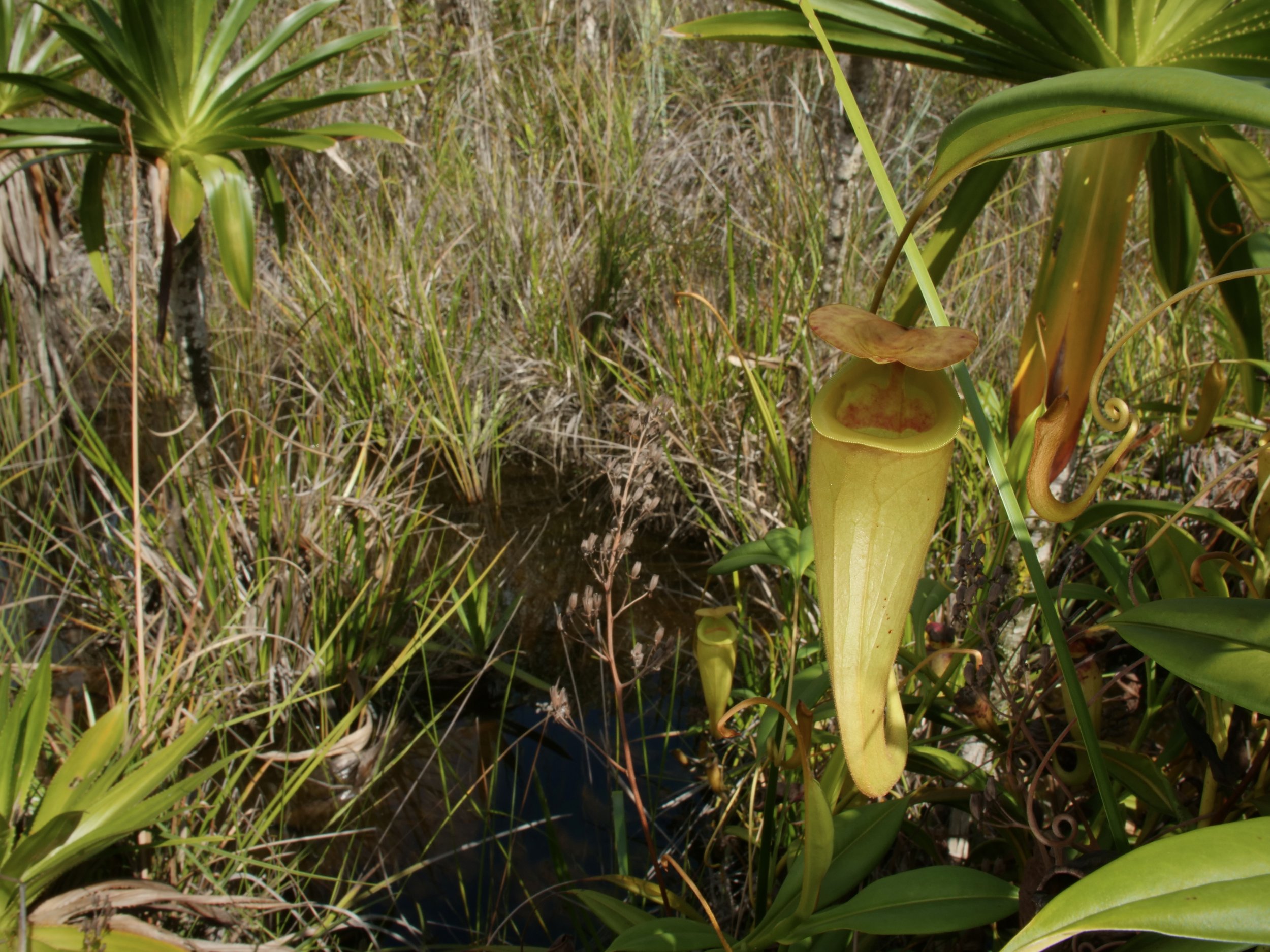
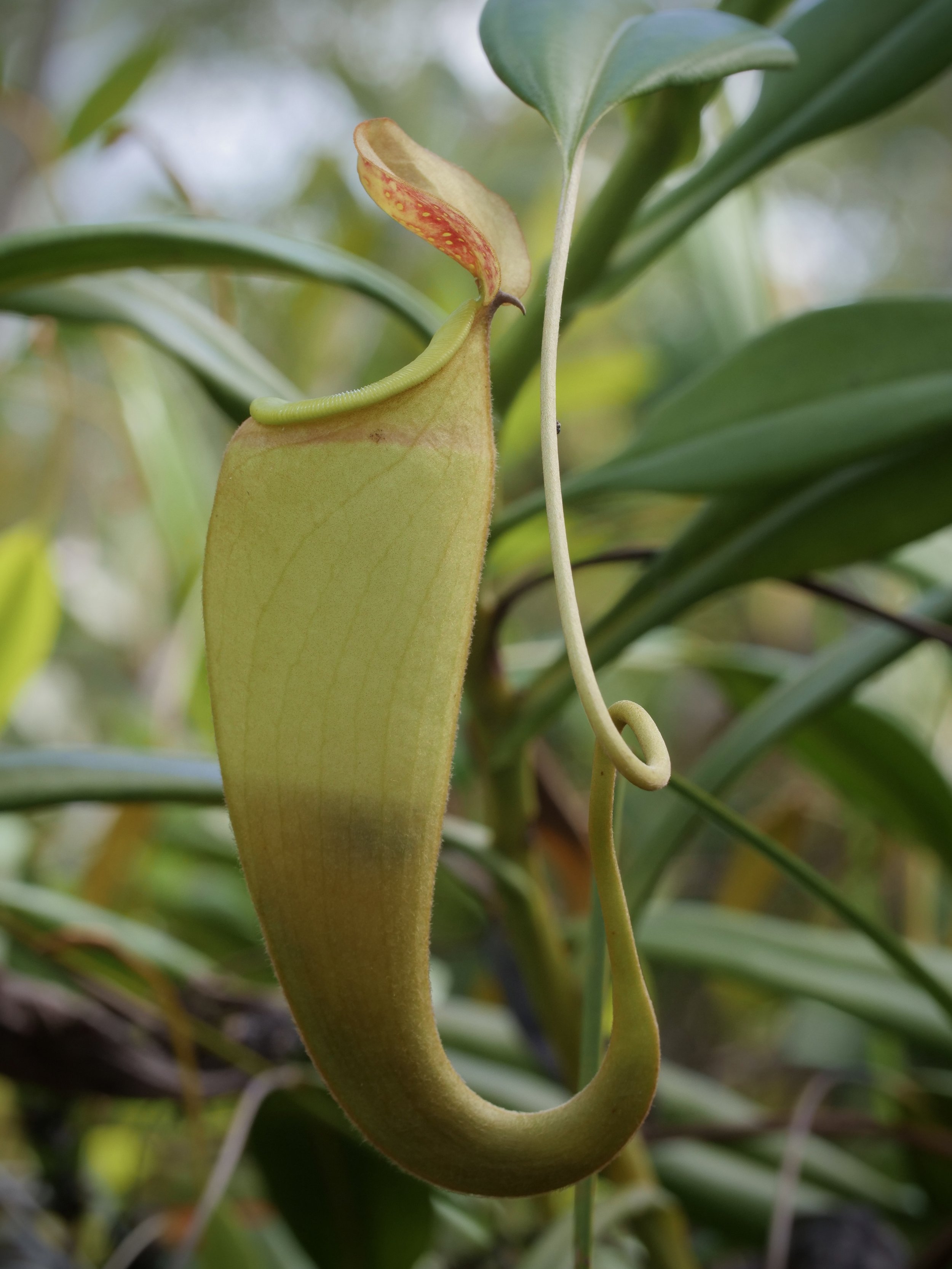
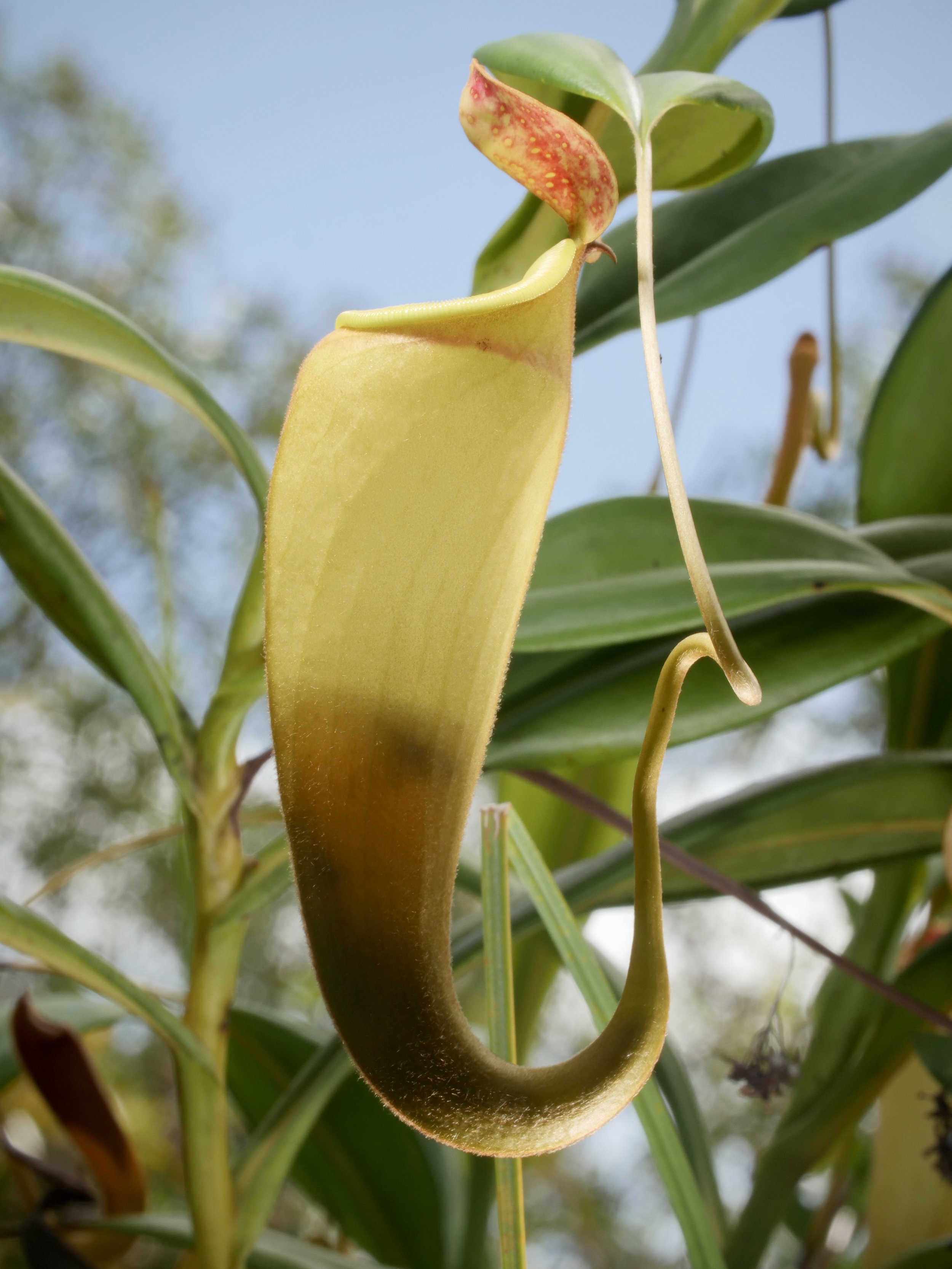




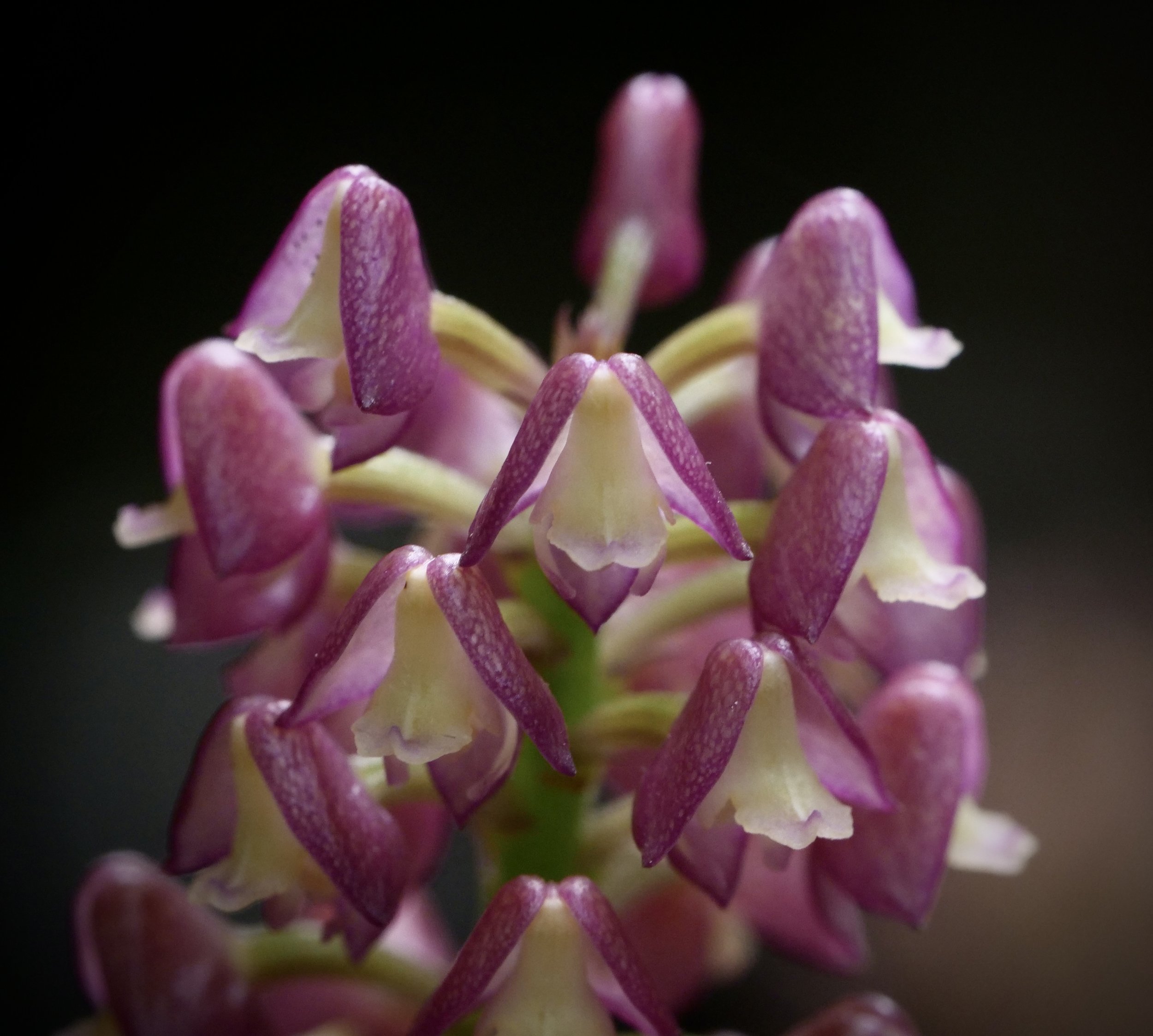
Pitcher plants Nepenthes madagascariensis, at Lake Ampitabe, Ankanin’ny Nofy, Madagascar
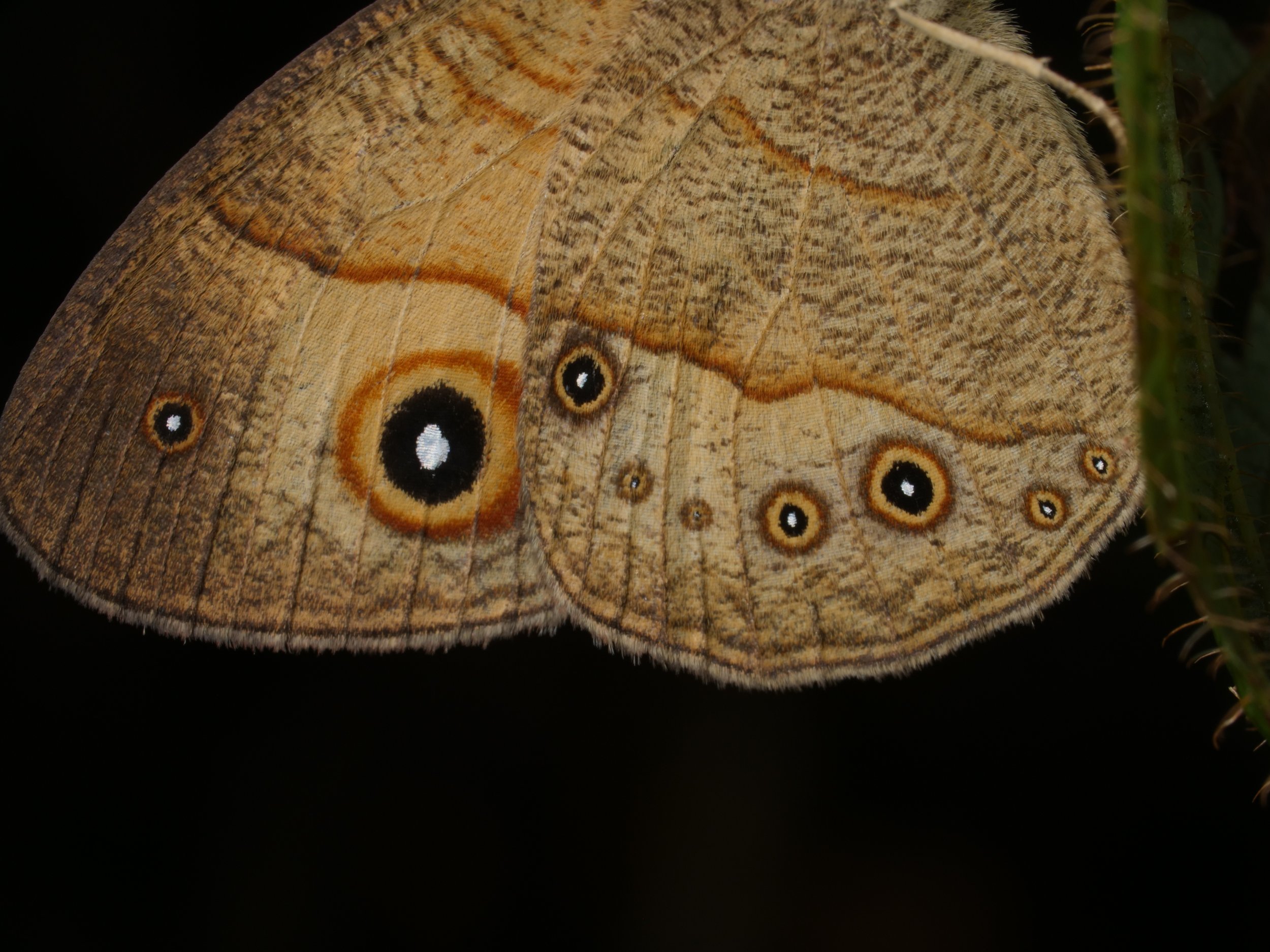
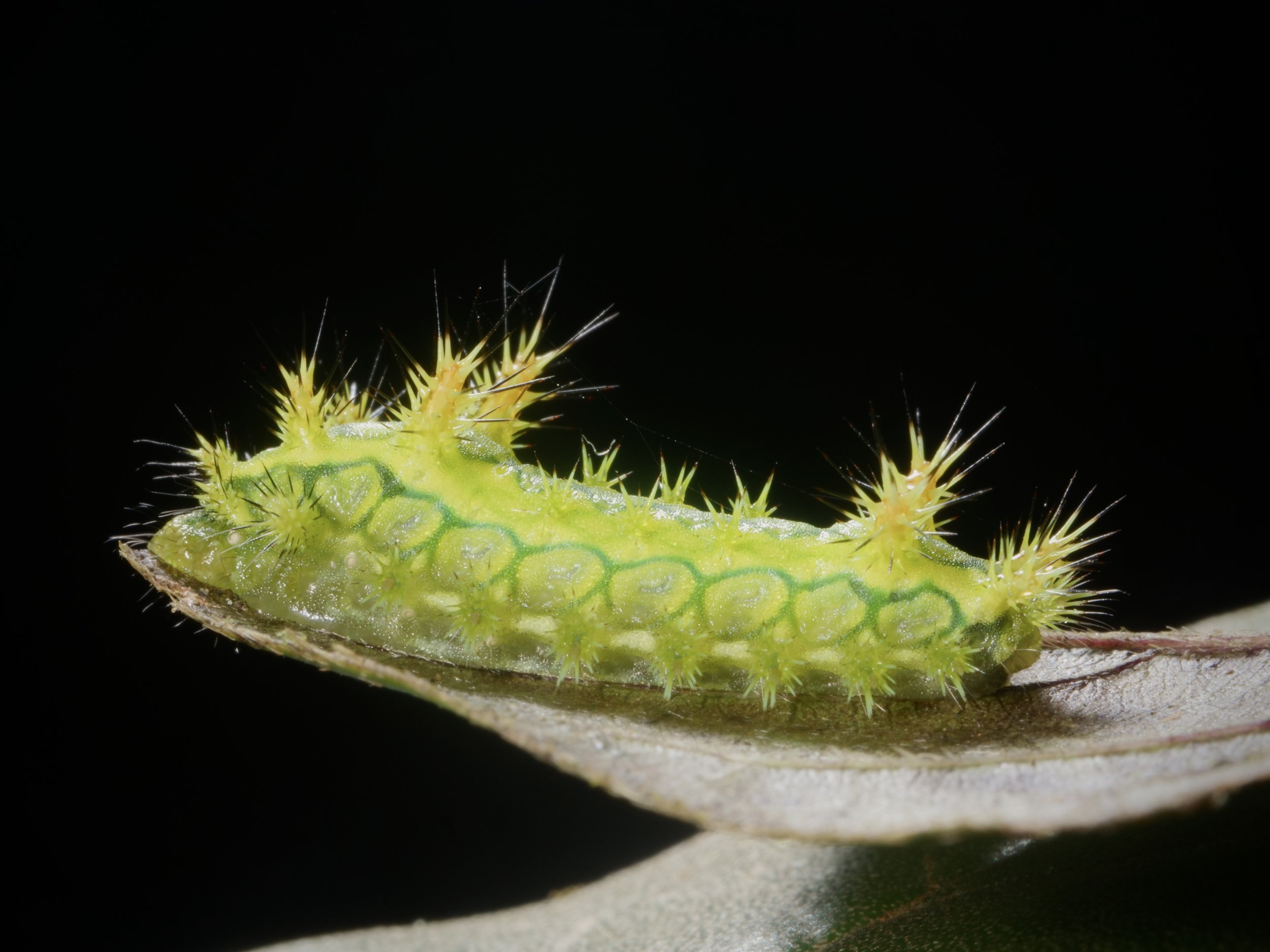
The ‘Galaxy’ caterpillar, a type of moth in the genus Eudocima, spotted on our trip in Madagascar. Those little blue spots and ‘false eye’ patterns are mesmerizing ✨ If you stare long enough you may get sucked into a back hole and vanish from the known universe…
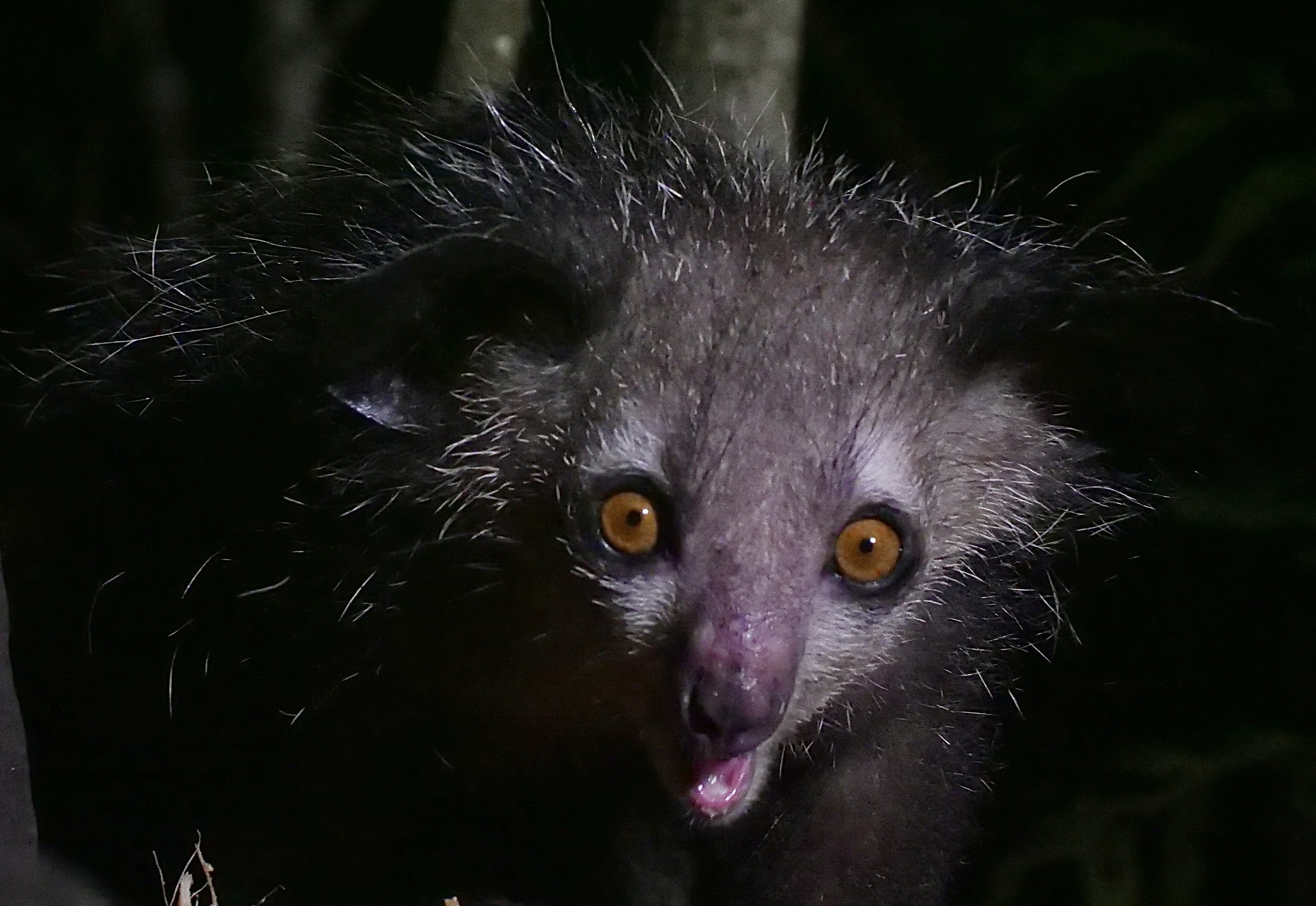
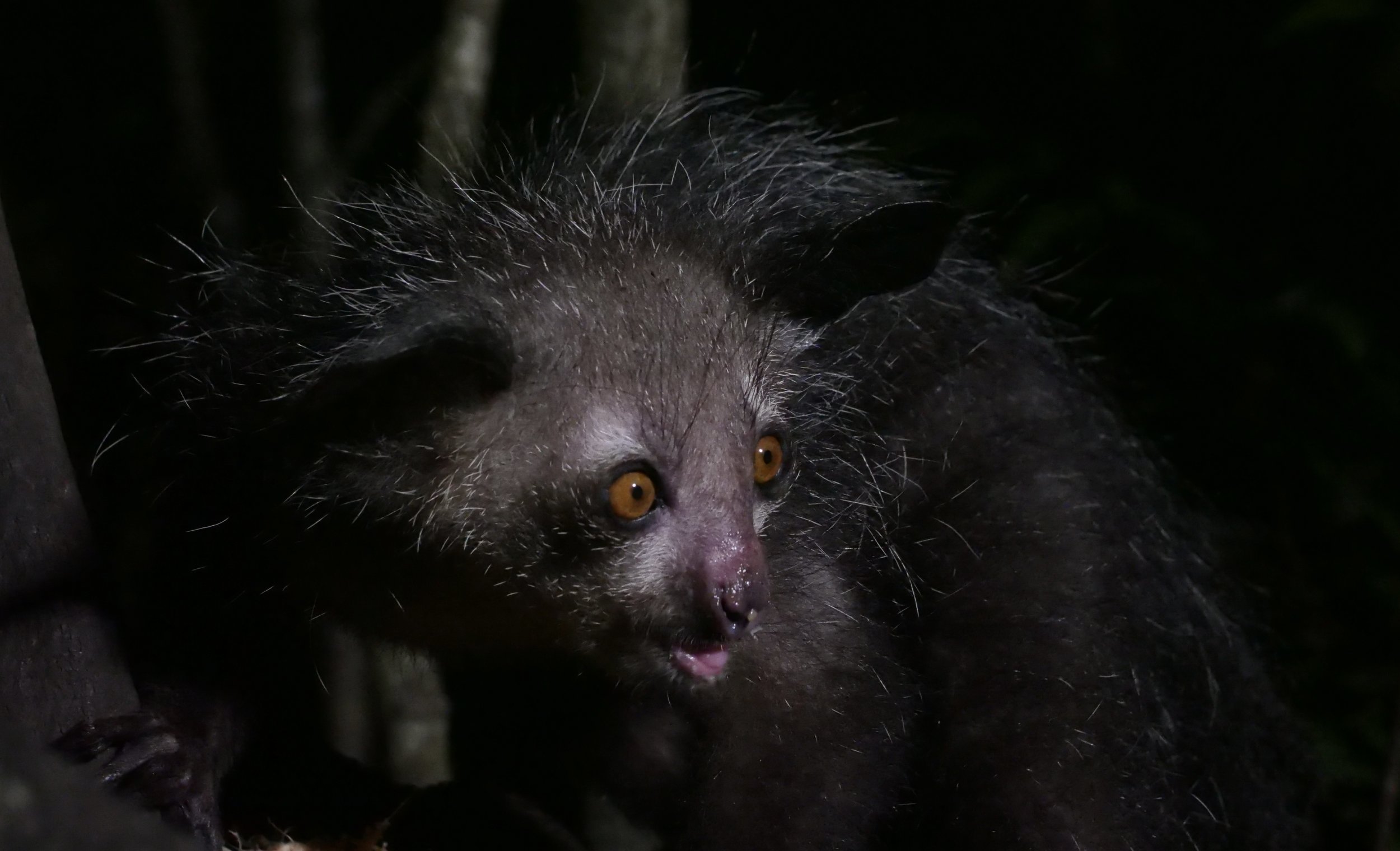
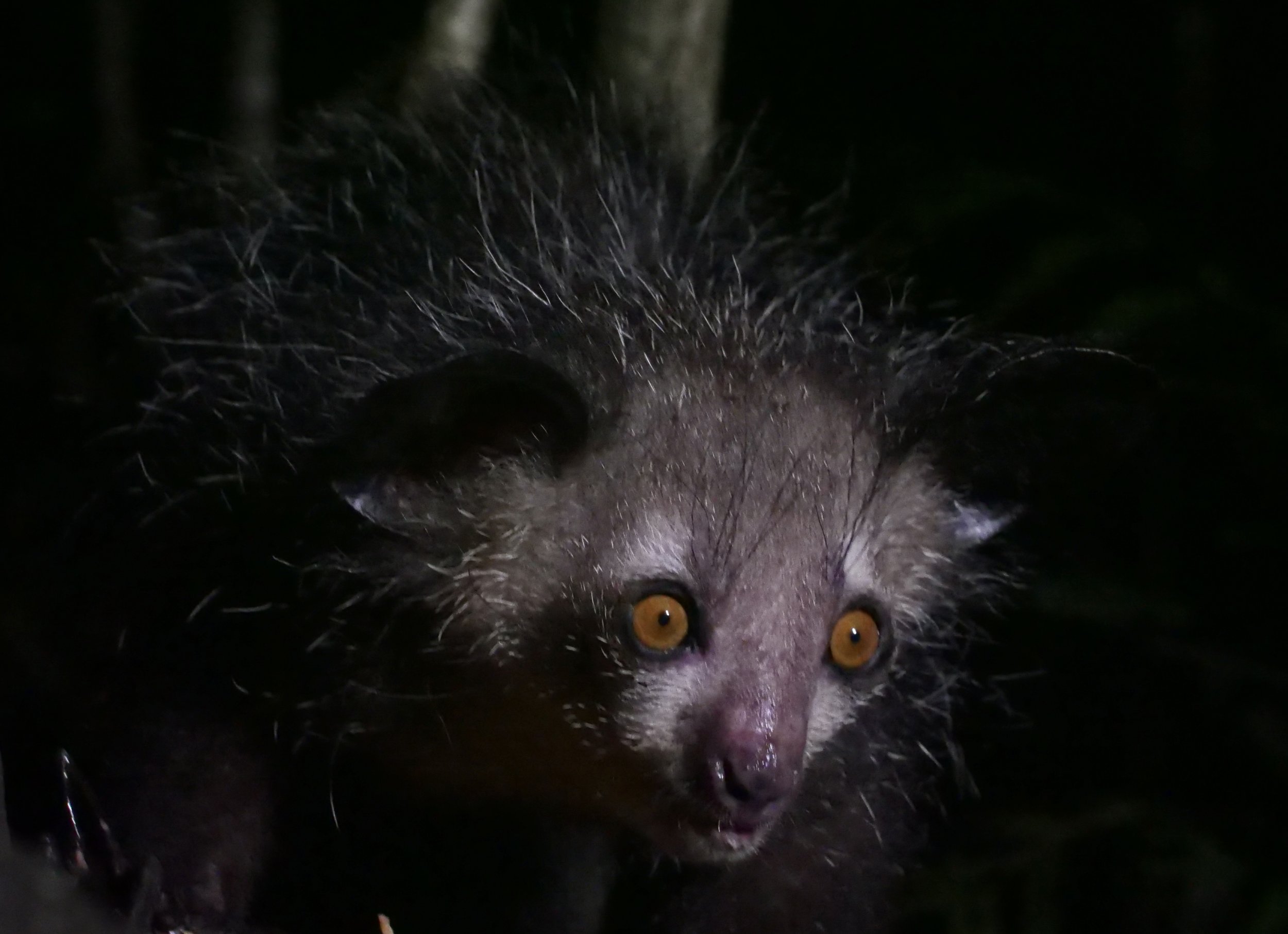
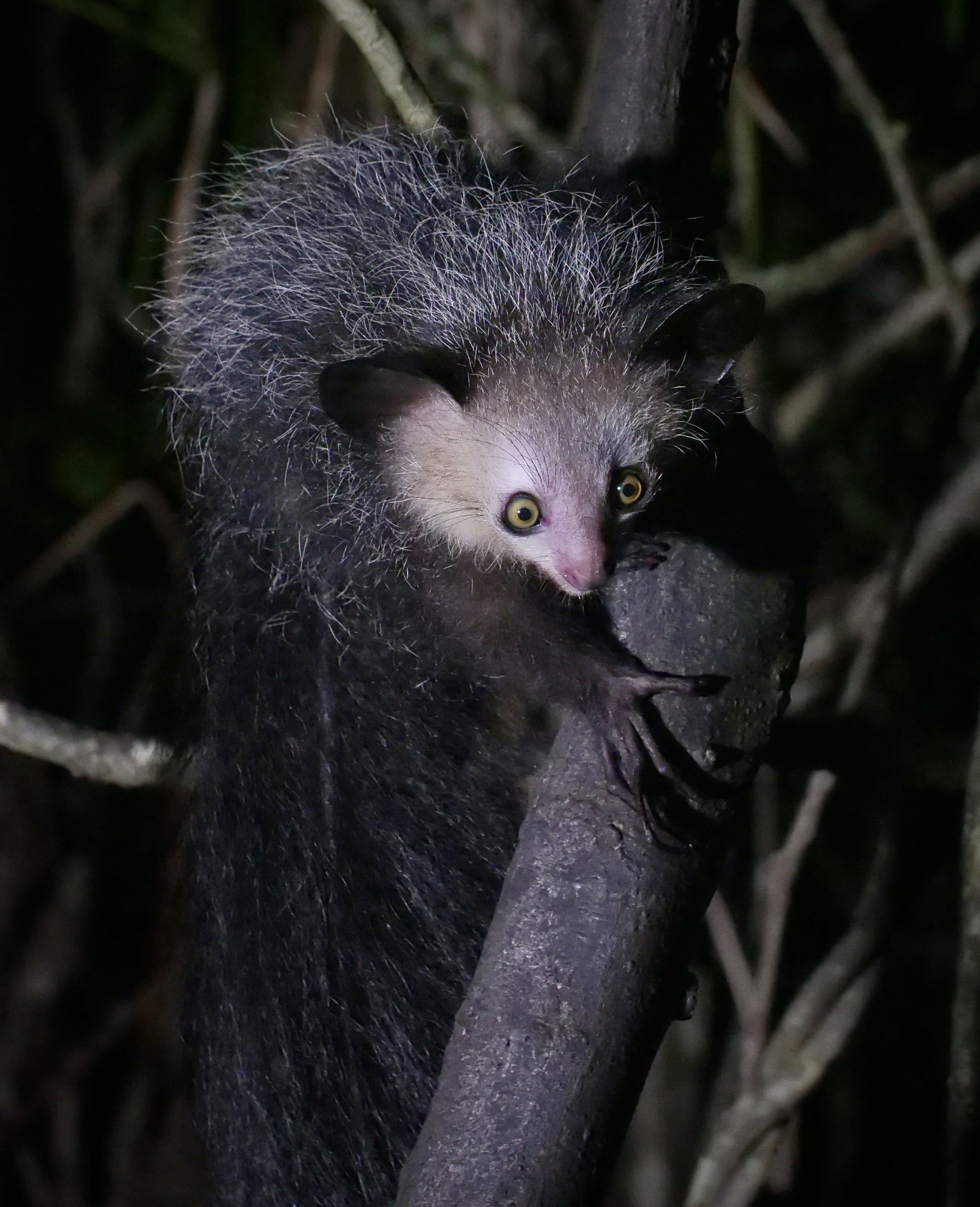
We spent three full days in the paradisiac Ankanin’ny Nofy doing jungle walks to photograph wildlife inlcuding frogs, chameleons, snakes, and lemurs. One of the nights, we visited an island to photograph the enigmatic Aye-Aye, one of the world’s strangest mammals.
The aye-aye Daubentonia madagascariensis is a long-fingered lemur, native to Madagascar with rodent-like teeth and a thin, elongated middle finger.
A once in a lifetime trip. Many thanks to our guides, Frank Pichardo and Jose Vieira at Tropical Herping, and Rainer Dolch, who made it an unforgettable wildlife experience.